Performs Latent Class Analysis (LCA) on binary response data using the Expectation-Maximization (EM) algorithm. LCA identifies unobserved (latent) subgroups of examinees with similar response patterns, and estimates both the class characteristics and individual membership probabilities.
Usage
LCA(
U,
ncls = 2,
na = NULL,
Z = NULL,
w = NULL,
maxiter = 100,
verbose = TRUE,
beta1 = 1,
beta2 = 1,
conf = NULL
)Arguments
- U
Either an object of class "exametrika" or raw data. When raw data is given, it is converted to the exametrika class with the
dataFormatfunction.- ncls
Number of latent classes to identify (between 2 and 20). Default is 2.
- na
Values to be treated as missing values.
- Z
Missing indicator matrix of type matrix or data.frame. Values of 1 indicate observed responses, while 0 indicates missing data.
- w
Item weight vector specifying the relative importance of each item.
- maxiter
Maximum number of EM algorithm iterations. Default is 100.
- verbose
Logical; if TRUE, displays progress during estimation. Default is TRUE.
- beta1
Beta distribution parameter 1 for prior density of class reference matrix. Default is 1.
- beta2
Beta distribution parameter 2 for prior density of class reference matrix. Default is 1.
- conf
Confirmatory IRP matrix (items x ncls) for test equating. Same format as the IRP output. Non-NA values are fixed throughout estimation, NA values are freely estimated. Fixed values must be in the open interval (0, 1). When row names are present, items are matched by label; otherwise by position. Default is NULL (fully exploratory).
Value
An object of class "exametrika" and "LCA" containing:
- msg
A character string indicating the model type.
- testlength
Length of the test (number of items).
- nobs
Sample size (number of rows in the dataset).
- Nclass
Number of latent classes specified.
- N_Cycle
Number of EM algorithm iterations performed.
- converge
Logical value indicating whether the algorithm converged within maxiter iterations
- TRP
Test Reference Profile vector showing expected scores for each latent class. Calculated as the column sum of the estimated class reference matrix.
- LCD
Latent Class Distribution vector showing the number of examinees assigned to each latent class.
- CMD
Class Membership Distribution vector showing the sum of membership probabilities for each latent class.
- Students
Class Membership Profile matrix showing the posterior probability of each examinee belonging to each latent class. The last column ("Estimate") indicates the most likely class assignment.
- IRP
Item Reference Profile matrix where each row represents an item and each column represents a latent class. Values indicate the probability of a correct response for members of that class.
- ItemFitIndices
Fit indices for each item. See also
ItemFit.- TestFitIndices
Overall fit indices for the test. See also
TestFit.
Details
Latent Class Analysis is a statistical method for identifying unobserved subgroups within a population based on observed response patterns. It assumes that examinees belong to one of several distinct latent classes, and that the probability of a correct response to each item depends on class membership.
The algorithm proceeds by:
Initializing class reference probabilities
Computing posterior class membership probabilities for each examinee (E-step)
Re-estimating class reference probabilities based on these memberships (M-step)
Iterating until convergence or reaching the maximum number of iterations
Unlike Item Response Theory (IRT), LCA treats latent variables as categorical rather than continuous, identifying distinct profiles rather than positions on a continuum.
References
Goodman, L. A. (1974). Exploratory latent structure analysis using both identifiable and unidentifiable models. Biometrika, 61(2), 215-231.
Lazarsfeld, P. F., & Henry, N. W. (1968). Latent structure analysis. Boston: Houghton Mifflin.
Examples
# \donttest{
# Fit a Latent Class Analysis model with 5 classes to the sample dataset
result.LCA <- LCA(J15S500, ncls = 5)
#>
iter 1 log_lik -3920.06
#>
iter 2 log_lik -3868.38
#>
iter 3 log_lik -3856.57
#>
iter 4 log_lik -3845.74
#>
iter 5 log_lik -3833.37
#>
iter 6 log_lik -3820.23
#>
iter 7 log_lik -3806.72
#>
iter 8 log_lik -3792.97
#>
iter 9 log_lik -3779.19
#>
iter 10 log_lik -3765.86
#>
iter 11 log_lik -3753.64
#>
iter 12 log_lik -3743.12
#>
iter 13 log_lik -3734.56
#>
iter 14 log_lik -3727.88
#>
iter 15 log_lik -3722.77
#>
iter 16 log_lik -3718.86
#>
iter 17 log_lik -3715.8
#>
iter 18 log_lik -3713.3
#>
iter 19 log_lik -3711.18
#>
iter 20 log_lik -3709.29
#>
iter 21 log_lik -3707.55
#>
iter 22 log_lik -3705.91
#>
iter 23 log_lik -3704.34
#>
iter 24 log_lik -3702.83
#>
iter 25 log_lik -3701.39
#>
iter 26 log_lik -3699.99
#>
iter 27 log_lik -3698.66
#>
iter 28 log_lik -3697.39
#>
iter 29 log_lik -3696.18
#>
iter 30 log_lik -3695.02
#>
iter 31 log_lik -3693.92
#>
iter 32 log_lik -3692.86
#>
iter 33 log_lik -3691.86
#>
iter 34 log_lik -3690.89
#>
iter 35 log_lik -3689.96
#>
iter 36 log_lik -3689.06
#>
iter 37 log_lik -3688.17
#>
iter 38 log_lik -3687.31
#>
iter 39 log_lik -3686.45
#>
iter 40 log_lik -3685.61
#>
iter 41 log_lik -3684.76
#>
iter 42 log_lik -3683.91
#>
iter 43 log_lik -3683.07
#>
iter 44 log_lik -3682.22
#>
iter 45 log_lik -3681.37
#>
iter 46 log_lik -3680.53
#>
iter 47 log_lik -3679.69
#>
iter 48 log_lik -3678.86
#>
iter 49 log_lik -3678.05
#>
iter 50 log_lik -3677.25
#>
iter 51 log_lik -3676.47
#>
iter 52 log_lik -3675.71
#>
iter 53 log_lik -3674.97
#>
iter 54 log_lik -3674.26
#>
iter 55 log_lik -3673.57
#>
iter 56 log_lik -3672.9
#>
iter 57 log_lik -3672.24
#>
iter 58 log_lik -3671.61
#>
iter 59 log_lik -3670.99
#>
iter 60 log_lik -3670.38
#>
iter 61 log_lik -3669.79
#>
iter 62 log_lik -3669.21
#>
iter 63 log_lik -3668.64
#>
iter 64 log_lik -3668.09
#>
iter 65 log_lik -3667.55
#>
iter 66 log_lik -3667.03
#>
iter 67 log_lik -3666.53
#>
iter 68 log_lik -3666.05
#>
iter 69 log_lik -3665.59
#>
iter 70 log_lik -3665.15
#>
iter 71 log_lik -3664.74
#>
iter 72 log_lik -3664.35
#>
iter 73 log_lik -3663.99
# Display the first few rows of student class membership probabilities
head(result.LCA$Students)
#> Membership 1 Membership 2 Membership 3 Membership 4 Membership 5
#> Student001 0.7839477684 0.171152798 0.004141844 4.075759e-02 3.744590e-12
#> Student002 0.0347378747 0.051502214 0.836022799 7.773694e-02 1.698776e-07
#> Student003 0.0146307878 0.105488644 0.801853496 3.343026e-02 4.459682e-02
#> Student004 0.0017251650 0.023436459 0.329648386 3.656488e-01 2.795412e-01
#> Student005 0.2133830569 0.784162066 0.001484616 2.492073e-08 9.702355e-04
#> Student006 0.0003846482 0.001141448 0.001288901 8.733869e-01 1.237981e-01
#> Estimate
#> Student001 1
#> Student002 3
#> Student003 3
#> Student004 4
#> Student005 2
#> Student006 4
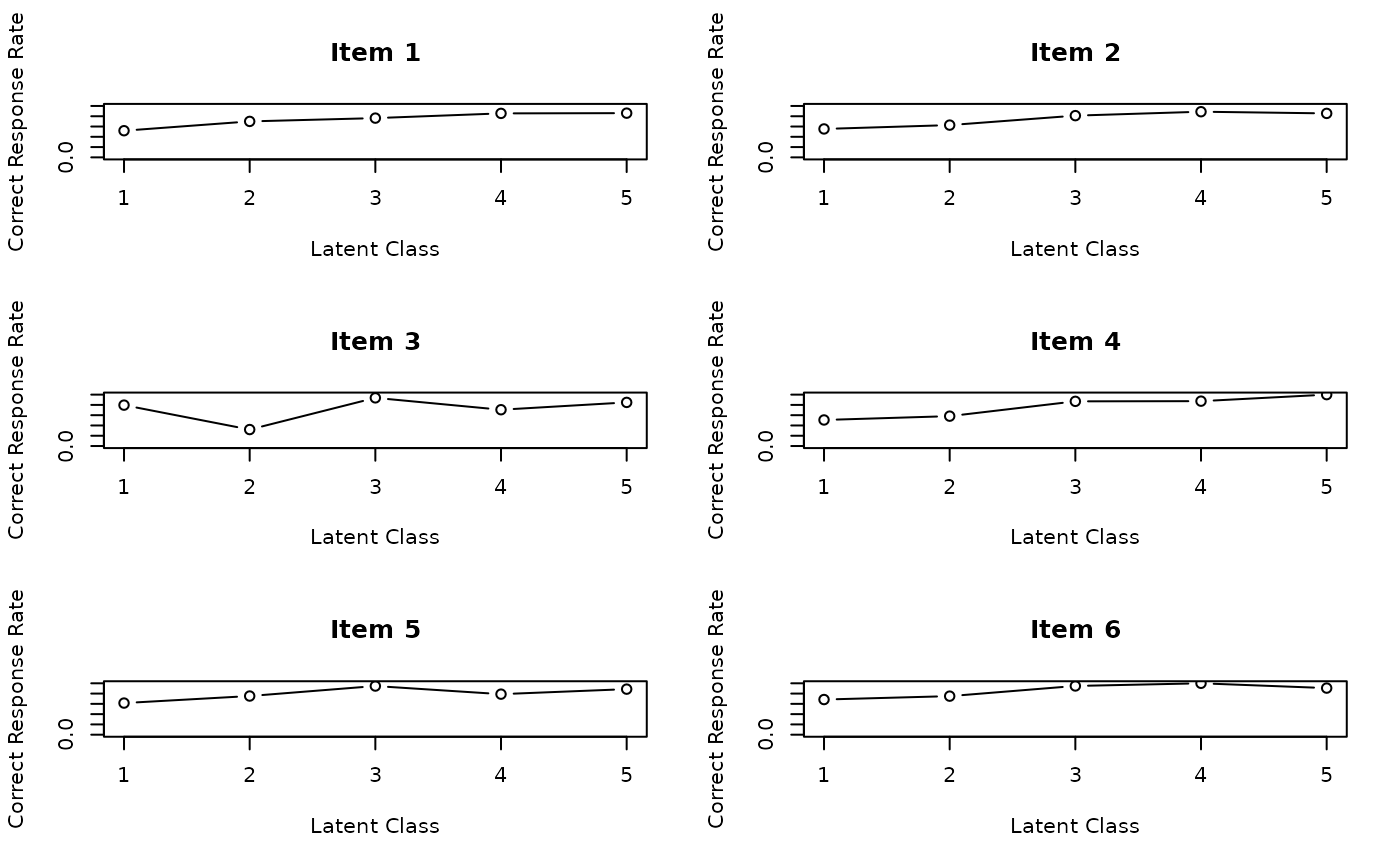
# Plot Item Response Profiles (IRP) for items 1-6 in a 2x3 grid
# Shows probability of correct response for each item across classes
plot(result.LCA, type = "IRP", items = 1:6, nc = 2, nr = 3)
 # Plot Class Membership Probabilities (CMP) for students 1-9 in a 3x3 grid
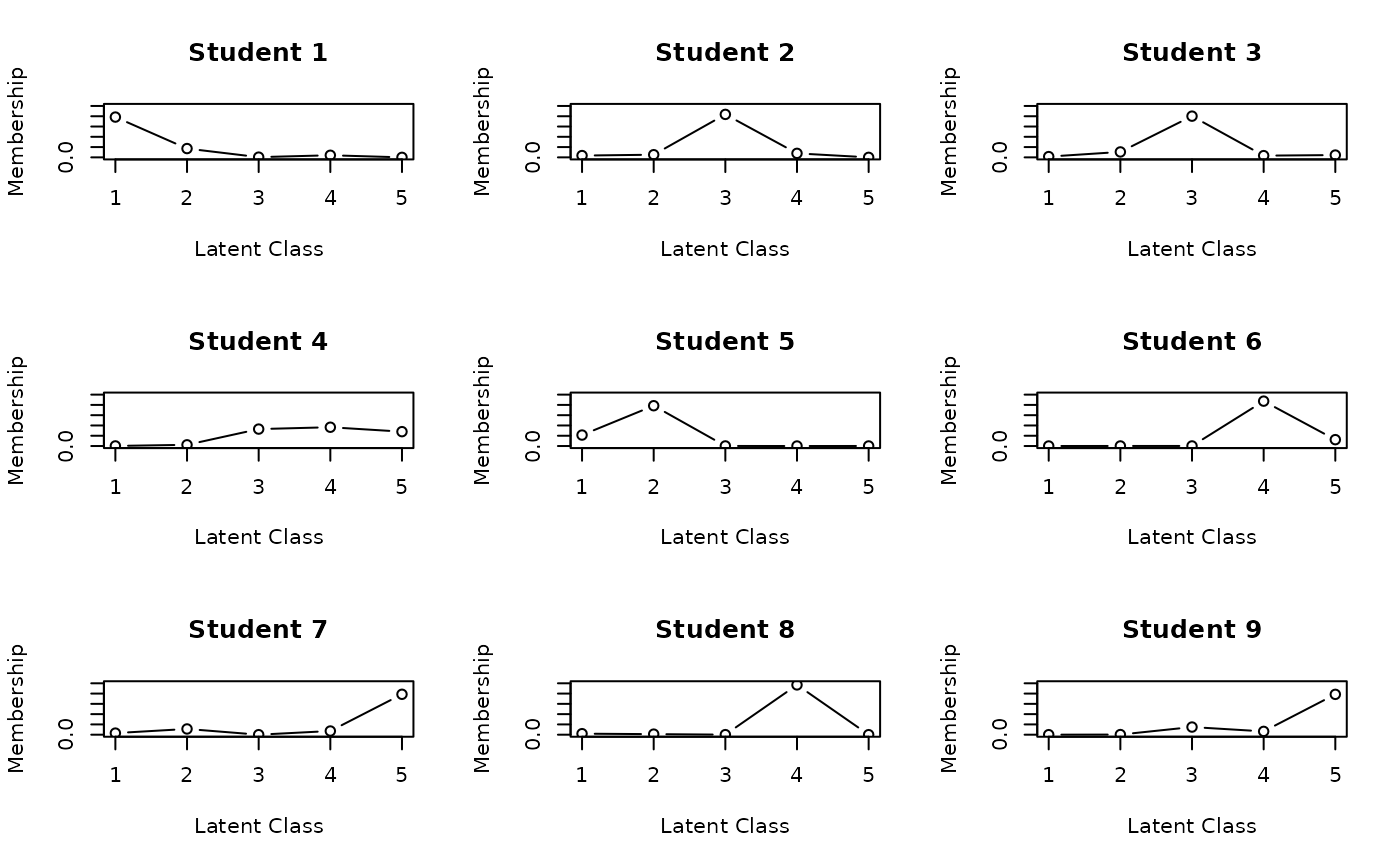
# Shows probability distribution of class membership for each student
plot(result.LCA, type = "CMP", students = 1:9, nc = 3, nr = 3)
# Plot Class Membership Probabilities (CMP) for students 1-9 in a 3x3 grid
# Shows probability distribution of class membership for each student
plot(result.LCA, type = "CMP", students = 1:9, nc = 3, nr = 3)
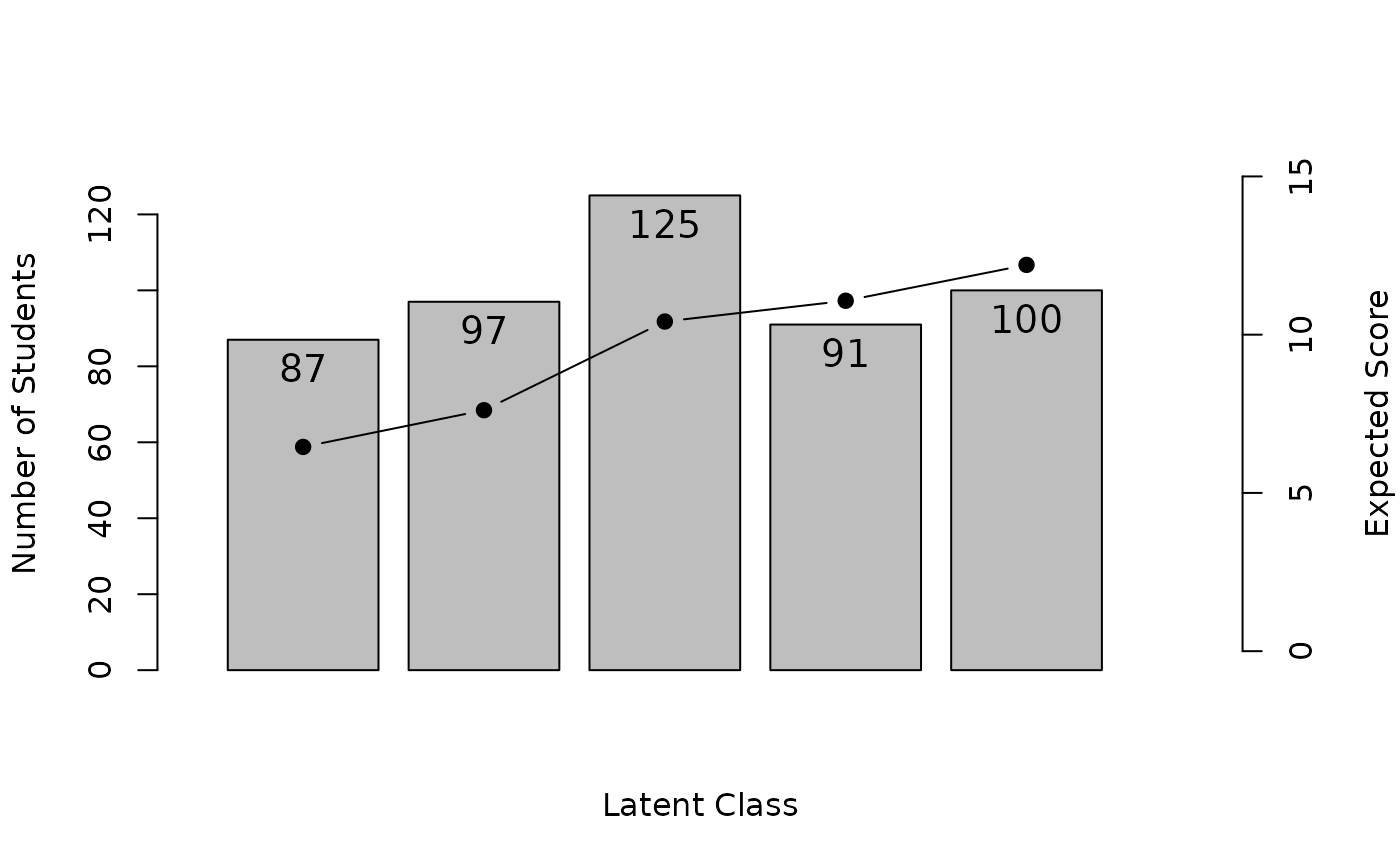
 # Plot Test Response Profile (TRP) showing expected scores for each class
plot(result.LCA, type = "TRP")
# Plot Test Response Profile (TRP) showing expected scores for each class
plot(result.LCA, type = "TRP")
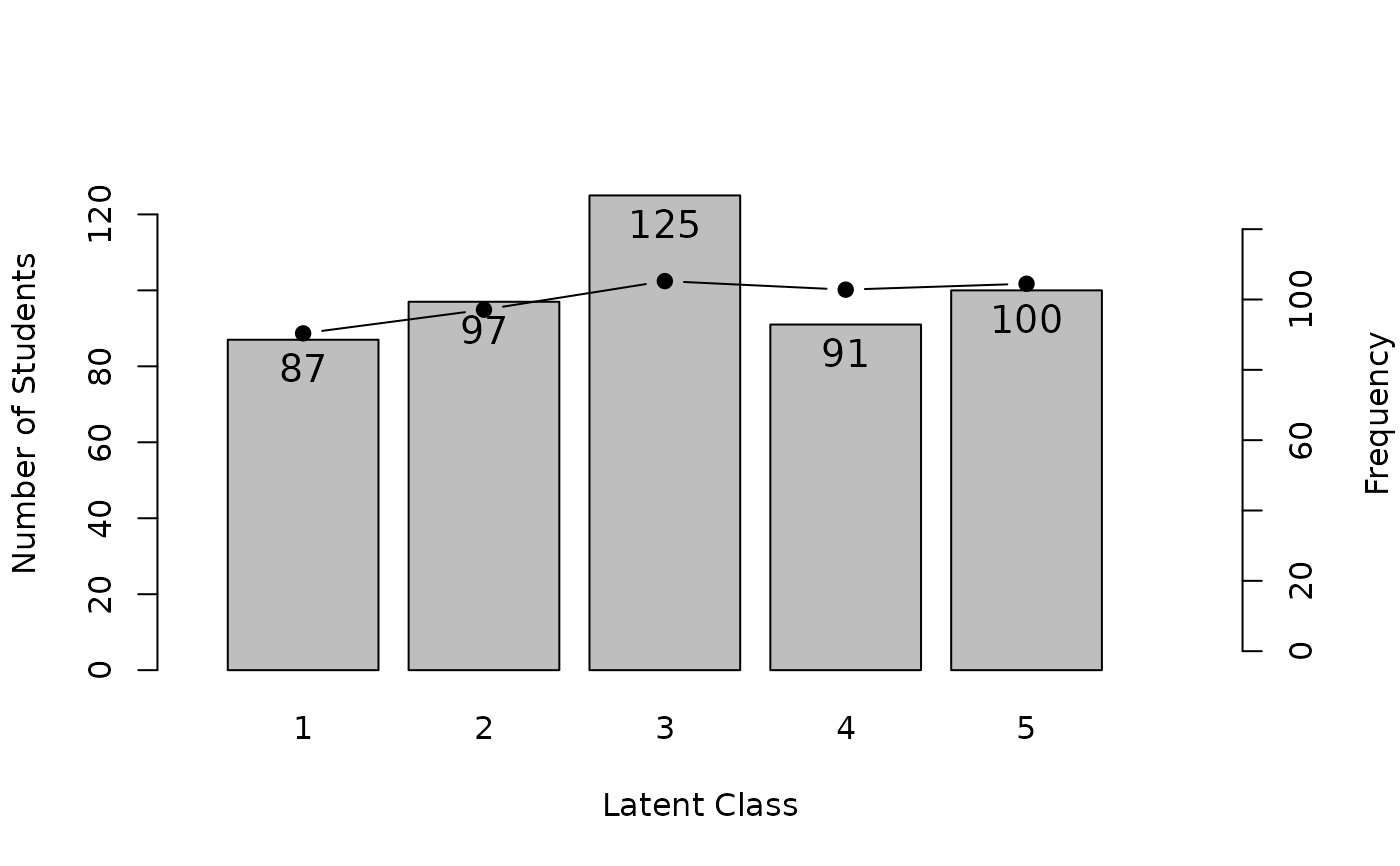
 # Plot Latent Class Distribution (LCD) showing class sizes
plot(result.LCA, type = "LCD")
# Plot Latent Class Distribution (LCD) showing class sizes
plot(result.LCA, type = "LCD")
 # Compare models with different numbers of classes
# (In practice, you might try more class counts)
lca2 <- LCA(J15S500, ncls = 2)
#>
iter 1 log_lik -4015.98
#>
iter 2 log_lik -3947.58
#>
iter 3 log_lik -3941.3
#>
iter 4 log_lik -3942.48
lca3 <- LCA(J15S500, ncls = 3)
#>
iter 1 log_lik -3955.4
#>
iter 2 log_lik -3904.63
#>
iter 3 log_lik -3890.82
#>
iter 4 log_lik -3880
#>
iter 5 log_lik -3870.82
#>
iter 6 log_lik -3863.52
#>
iter 7 log_lik -3857.89
#>
iter 8 log_lik -3853.58
#>
iter 9 log_lik -3850.31
#>
iter 10 log_lik -3847.86
#>
iter 11 log_lik -3846.05
#>
iter 12 log_lik -3844.72
#>
iter 13 log_lik -3843.74
#>
iter 14 log_lik -3843.02
#>
iter 15 log_lik -3842.48
#>
iter 16 log_lik -3842.07
#>
iter 17 log_lik -3841.76
lca4 <- LCA(J15S500, ncls = 4)
#>
iter 1 log_lik -3931.94
#>
iter 2 log_lik -3881.61
#>
iter 3 log_lik -3871.87
#>
iter 4 log_lik -3863.93
#>
iter 5 log_lik -3854.73
#>
iter 6 log_lik -3845.5
#>
iter 7 log_lik -3837.24
#>
iter 8 log_lik -3830.25
#>
iter 9 log_lik -3824.31
#>
iter 10 log_lik -3819.07
#>
iter 11 log_lik -3814.18
#>
iter 12 log_lik -3809.38
#>
iter 13 log_lik -3804.51
#>
iter 14 log_lik -3799.5
#>
iter 15 log_lik -3794.39
#>
iter 16 log_lik -3789.28
#>
iter 17 log_lik -3784.29
#>
iter 18 log_lik -3779.54
#>
iter 19 log_lik -3775.13
#>
iter 20 log_lik -3771.13
#>
iter 21 log_lik -3767.57
#>
iter 22 log_lik -3764.45
#>
iter 23 log_lik -3761.76
#>
iter 24 log_lik -3759.45
#>
iter 25 log_lik -3757.48
#>
iter 26 log_lik -3755.81
#>
iter 27 log_lik -3754.39
#>
iter 28 log_lik -3753.18
#>
iter 29 log_lik -3752.15
#>
iter 30 log_lik -3751.26
#>
iter 31 log_lik -3750.49
#>
iter 32 log_lik -3749.82
#>
iter 33 log_lik -3749.23
#>
iter 34 log_lik -3748.72
#>
iter 35 log_lik -3748.26
#>
iter 36 log_lik -3747.85
#>
iter 37 log_lik -3747.48
lca5 <- LCA(J15S500, ncls = 5)
#>
iter 1 log_lik -3920.06
#>
iter 2 log_lik -3868.38
#>
iter 3 log_lik -3856.57
#>
iter 4 log_lik -3845.74
#>
iter 5 log_lik -3833.37
#>
iter 6 log_lik -3820.23
#>
iter 7 log_lik -3806.72
#>
iter 8 log_lik -3792.97
#>
iter 9 log_lik -3779.19
#>
iter 10 log_lik -3765.86
#>
iter 11 log_lik -3753.64
#>
iter 12 log_lik -3743.12
#>
iter 13 log_lik -3734.56
#>
iter 14 log_lik -3727.88
#>
iter 15 log_lik -3722.77
#>
iter 16 log_lik -3718.86
#>
iter 17 log_lik -3715.8
#>
iter 18 log_lik -3713.3
#>
iter 19 log_lik -3711.18
#>
iter 20 log_lik -3709.29
#>
iter 21 log_lik -3707.55
#>
iter 22 log_lik -3705.91
#>
iter 23 log_lik -3704.34
#>
iter 24 log_lik -3702.83
#>
iter 25 log_lik -3701.39
#>
iter 26 log_lik -3699.99
#>
iter 27 log_lik -3698.66
#>
iter 28 log_lik -3697.39
#>
iter 29 log_lik -3696.18
#>
iter 30 log_lik -3695.02
#>
iter 31 log_lik -3693.92
#>
iter 32 log_lik -3692.86
#>
iter 33 log_lik -3691.86
#>
iter 34 log_lik -3690.89
#>
iter 35 log_lik -3689.96
#>
iter 36 log_lik -3689.06
#>
iter 37 log_lik -3688.17
#>
iter 38 log_lik -3687.31
#>
iter 39 log_lik -3686.45
#>
iter 40 log_lik -3685.61
#>
iter 41 log_lik -3684.76
#>
iter 42 log_lik -3683.91
#>
iter 43 log_lik -3683.07
#>
iter 44 log_lik -3682.22
#>
iter 45 log_lik -3681.37
#>
iter 46 log_lik -3680.53
#>
iter 47 log_lik -3679.69
#>
iter 48 log_lik -3678.86
#>
iter 49 log_lik -3678.05
#>
iter 50 log_lik -3677.25
#>
iter 51 log_lik -3676.47
#>
iter 52 log_lik -3675.71
#>
iter 53 log_lik -3674.97
#>
iter 54 log_lik -3674.26
#>
iter 55 log_lik -3673.57
#>
iter 56 log_lik -3672.9
#>
iter 57 log_lik -3672.24
#>
iter 58 log_lik -3671.61
#>
iter 59 log_lik -3670.99
#>
iter 60 log_lik -3670.38
#>
iter 61 log_lik -3669.79
#>
iter 62 log_lik -3669.21
#>
iter 63 log_lik -3668.64
#>
iter 64 log_lik -3668.09
#>
iter 65 log_lik -3667.55
#>
iter 66 log_lik -3667.03
#>
iter 67 log_lik -3666.53
#>
iter 68 log_lik -3666.05
#>
iter 69 log_lik -3665.59
#>
iter 70 log_lik -3665.15
#>
iter 71 log_lik -3664.74
#>
iter 72 log_lik -3664.35
#>
iter 73 log_lik -3663.99
# Compare BIC values to select optimal number of classes
# (Lower BIC indicates better fit)
data.frame(
Classes = 2:5,
BIC = c(
lca2$TestFitIndices$BIC,
lca3$TestFitIndices$BIC,
lca4$TestFitIndices$BIC,
lca5$TestFitIndices$BIC
)
)
#> Classes BIC
#> 1 2 -349.9323
#> 2 3 -461.9099
#> 3 4 -557.2459
#> 4 5 -630.9953
# }
# Compare models with different numbers of classes
# (In practice, you might try more class counts)
lca2 <- LCA(J15S500, ncls = 2)
#>
iter 1 log_lik -4015.98
#>
iter 2 log_lik -3947.58
#>
iter 3 log_lik -3941.3
#>
iter 4 log_lik -3942.48
lca3 <- LCA(J15S500, ncls = 3)
#>
iter 1 log_lik -3955.4
#>
iter 2 log_lik -3904.63
#>
iter 3 log_lik -3890.82
#>
iter 4 log_lik -3880
#>
iter 5 log_lik -3870.82
#>
iter 6 log_lik -3863.52
#>
iter 7 log_lik -3857.89
#>
iter 8 log_lik -3853.58
#>
iter 9 log_lik -3850.31
#>
iter 10 log_lik -3847.86
#>
iter 11 log_lik -3846.05
#>
iter 12 log_lik -3844.72
#>
iter 13 log_lik -3843.74
#>
iter 14 log_lik -3843.02
#>
iter 15 log_lik -3842.48
#>
iter 16 log_lik -3842.07
#>
iter 17 log_lik -3841.76
lca4 <- LCA(J15S500, ncls = 4)
#>
iter 1 log_lik -3931.94
#>
iter 2 log_lik -3881.61
#>
iter 3 log_lik -3871.87
#>
iter 4 log_lik -3863.93
#>
iter 5 log_lik -3854.73
#>
iter 6 log_lik -3845.5
#>
iter 7 log_lik -3837.24
#>
iter 8 log_lik -3830.25
#>
iter 9 log_lik -3824.31
#>
iter 10 log_lik -3819.07
#>
iter 11 log_lik -3814.18
#>
iter 12 log_lik -3809.38
#>
iter 13 log_lik -3804.51
#>
iter 14 log_lik -3799.5
#>
iter 15 log_lik -3794.39
#>
iter 16 log_lik -3789.28
#>
iter 17 log_lik -3784.29
#>
iter 18 log_lik -3779.54
#>
iter 19 log_lik -3775.13
#>
iter 20 log_lik -3771.13
#>
iter 21 log_lik -3767.57
#>
iter 22 log_lik -3764.45
#>
iter 23 log_lik -3761.76
#>
iter 24 log_lik -3759.45
#>
iter 25 log_lik -3757.48
#>
iter 26 log_lik -3755.81
#>
iter 27 log_lik -3754.39
#>
iter 28 log_lik -3753.18
#>
iter 29 log_lik -3752.15
#>
iter 30 log_lik -3751.26
#>
iter 31 log_lik -3750.49
#>
iter 32 log_lik -3749.82
#>
iter 33 log_lik -3749.23
#>
iter 34 log_lik -3748.72
#>
iter 35 log_lik -3748.26
#>
iter 36 log_lik -3747.85
#>
iter 37 log_lik -3747.48
lca5 <- LCA(J15S500, ncls = 5)
#>
iter 1 log_lik -3920.06
#>
iter 2 log_lik -3868.38
#>
iter 3 log_lik -3856.57
#>
iter 4 log_lik -3845.74
#>
iter 5 log_lik -3833.37
#>
iter 6 log_lik -3820.23
#>
iter 7 log_lik -3806.72
#>
iter 8 log_lik -3792.97
#>
iter 9 log_lik -3779.19
#>
iter 10 log_lik -3765.86
#>
iter 11 log_lik -3753.64
#>
iter 12 log_lik -3743.12
#>
iter 13 log_lik -3734.56
#>
iter 14 log_lik -3727.88
#>
iter 15 log_lik -3722.77
#>
iter 16 log_lik -3718.86
#>
iter 17 log_lik -3715.8
#>
iter 18 log_lik -3713.3
#>
iter 19 log_lik -3711.18
#>
iter 20 log_lik -3709.29
#>
iter 21 log_lik -3707.55
#>
iter 22 log_lik -3705.91
#>
iter 23 log_lik -3704.34
#>
iter 24 log_lik -3702.83
#>
iter 25 log_lik -3701.39
#>
iter 26 log_lik -3699.99
#>
iter 27 log_lik -3698.66
#>
iter 28 log_lik -3697.39
#>
iter 29 log_lik -3696.18
#>
iter 30 log_lik -3695.02
#>
iter 31 log_lik -3693.92
#>
iter 32 log_lik -3692.86
#>
iter 33 log_lik -3691.86
#>
iter 34 log_lik -3690.89
#>
iter 35 log_lik -3689.96
#>
iter 36 log_lik -3689.06
#>
iter 37 log_lik -3688.17
#>
iter 38 log_lik -3687.31
#>
iter 39 log_lik -3686.45
#>
iter 40 log_lik -3685.61
#>
iter 41 log_lik -3684.76
#>
iter 42 log_lik -3683.91
#>
iter 43 log_lik -3683.07
#>
iter 44 log_lik -3682.22
#>
iter 45 log_lik -3681.37
#>
iter 46 log_lik -3680.53
#>
iter 47 log_lik -3679.69
#>
iter 48 log_lik -3678.86
#>
iter 49 log_lik -3678.05
#>
iter 50 log_lik -3677.25
#>
iter 51 log_lik -3676.47
#>
iter 52 log_lik -3675.71
#>
iter 53 log_lik -3674.97
#>
iter 54 log_lik -3674.26
#>
iter 55 log_lik -3673.57
#>
iter 56 log_lik -3672.9
#>
iter 57 log_lik -3672.24
#>
iter 58 log_lik -3671.61
#>
iter 59 log_lik -3670.99
#>
iter 60 log_lik -3670.38
#>
iter 61 log_lik -3669.79
#>
iter 62 log_lik -3669.21
#>
iter 63 log_lik -3668.64
#>
iter 64 log_lik -3668.09
#>
iter 65 log_lik -3667.55
#>
iter 66 log_lik -3667.03
#>
iter 67 log_lik -3666.53
#>
iter 68 log_lik -3666.05
#>
iter 69 log_lik -3665.59
#>
iter 70 log_lik -3665.15
#>
iter 71 log_lik -3664.74
#>
iter 72 log_lik -3664.35
#>
iter 73 log_lik -3663.99
# Compare BIC values to select optimal number of classes
# (Lower BIC indicates better fit)
data.frame(
Classes = 2:5,
BIC = c(
lca2$TestFitIndices$BIC,
lca3$TestFitIndices$BIC,
lca4$TestFitIndices$BIC,
lca5$TestFitIndices$BIC
)
)
#> Classes BIC
#> 1 2 -349.9323
#> 2 3 -461.9099
#> 3 4 -557.2459
#> 4 5 -630.9953
# }
