Note: Some computationally intensive examples below are shown with
eval=FALSEto keep CRAN build times short. For full rendered output, see the pkgdown site.
Biclustering and Ranklustering
Biclustering and Ranklustering simultaneously cluster items into
fields and examinees into classes/ranks. The difference is specified via
the method option:
-
method = "B": Biclustering (no filtering matrix) -
method = "R": Ranklustering (with filtering matrix for ordered ranks)
Biclustering
Biclustering(J35S515, nfld = 5, ncls = 6, method = "B")
#> Biclustering Analysis
#>
#> Biclustering Reference Matrix Profile
#> Class1 Class2 Class3 Class4 Class5 Class6
#> Field1 0.6236 0.8636 0.8718 0.898 0.952 1.000
#> Field2 0.0627 0.3332 0.4255 0.919 0.990 1.000
#> Field3 0.2008 0.5431 0.2281 0.475 0.706 1.000
#> Field4 0.0495 0.2455 0.0782 0.233 0.648 0.983
#> Field5 0.0225 0.0545 0.0284 0.043 0.160 0.983
#>
#> Field Reference Profile Indices
#> Alpha A Beta B Gamma C
#> Field1 1 0.240 1 0.624 0.0 0.0000
#> Field2 3 0.493 3 0.426 0.0 0.0000
#> Field3 1 0.342 4 0.475 0.2 -0.3149
#> Field4 4 0.415 5 0.648 0.2 -0.1673
#> Field5 5 0.823 5 0.160 0.2 -0.0261
#>
#> Class 1 Class 2 Class 3 Class 4 Class 5 Class 6
#> Test Reference Profile 4.431 11.894 8.598 16.002 23.326 34.713
#> Latent Class Ditribution 157.000 64.000 82.000 106.000 89.000 17.000
#> Class Membership Distribution 146.105 73.232 85.753 106.414 86.529 16.968
#>
#> Field Membership Profile
#> CRR LFE Field1 Field2 Field3 Field4 Field5
#> Item01 0.850 1 1.000 0.000 0.000 0.000 0.000
#> Item31 0.812 1 1.000 0.000 0.000 0.000 0.000
#> Item32 0.808 1 1.000 0.000 0.000 0.000 0.000
#> Item21 0.616 2 0.000 1.000 0.000 0.000 0.000
#> Item23 0.600 2 0.000 1.000 0.000 0.000 0.000
#> Item22 0.586 2 0.000 1.000 0.000 0.000 0.000
#> Item24 0.567 2 0.000 1.000 0.000 0.000 0.000
#> Item25 0.491 2 0.000 1.000 0.000 0.000 0.000
#> Item11 0.476 2 0.000 1.000 0.000 0.000 0.000
#> Item26 0.452 2 0.000 1.000 0.000 0.000 0.000
#> Item27 0.414 2 0.000 1.000 0.000 0.000 0.000
#> Item07 0.573 3 0.000 0.000 1.000 0.000 0.000
#> Item03 0.458 3 0.000 0.000 1.000 0.000 0.000
#> Item33 0.437 3 0.000 0.000 1.000 0.000 0.000
#> Item02 0.392 3 0.000 0.000 1.000 0.000 0.000
#> Item09 0.390 3 0.000 0.000 1.000 0.000 0.000
#> Item10 0.353 3 0.000 0.000 1.000 0.000 0.000
#> Item08 0.350 3 0.000 0.000 1.000 0.000 0.000
#> Item12 0.340 4 0.000 0.000 0.000 1.000 0.000
#> Item04 0.303 4 0.000 0.000 0.000 1.000 0.000
#> Item17 0.276 4 0.000 0.000 0.000 1.000 0.000
#> Item05 0.250 4 0.000 0.000 0.000 1.000 0.000
#> Item13 0.237 4 0.000 0.000 0.000 1.000 0.000
#> Item34 0.229 4 0.000 0.000 0.000 1.000 0.000
#> Item29 0.227 4 0.000 0.000 0.000 1.000 0.000
#> Item28 0.221 4 0.000 0.000 0.000 1.000 0.000
#> Item06 0.216 4 0.000 0.000 0.000 1.000 0.000
#> Item16 0.216 4 0.000 0.000 0.000 1.000 0.000
#> Item35 0.155 5 0.000 0.000 0.000 0.000 1.000
#> Item14 0.126 5 0.000 0.000 0.000 0.000 1.000
#> Item15 0.087 5 0.000 0.000 0.000 0.000 1.000
#> Item30 0.085 5 0.000 0.000 0.000 0.000 1.000
#> Item20 0.054 5 0.000 0.000 0.000 0.000 1.000
#> Item19 0.052 5 0.000 0.000 0.000 0.000 1.000
#> Item18 0.049 5 0.000 0.000 0.000 0.000 1.000
#> Latent Field Distribution
#> Field 1 Field 2 Field 3 Field 4 Field 5
#> N of Items 3 8 7 10 7
#>
#> Model Fit Indices
#> Number of Latent Class : 6
#> Number of Latent Field: 5
#> Number of EM cycle: 33
#> value
#> model_log_like -6884.582
#> bench_log_like -5891.314
#> null_log_like -9862.114
#> model_Chi_sq 1986.535
#> null_Chi_sq 7941.601
#> model_df 1160.000
#> null_df 1155.000
#> NFI 0.750
#> RFI 0.751
#> IFI 0.878
#> TLI 0.879
#> CFI 0.878
#> RMSEA 0.037
#> AIC -333.465
#> CAIC -6416.699
#> BIC -5256.699Ranklustering
result.Ranklustering <- Biclustering(J35S515, nfld = 5, ncls = 6, method = "R")
plot(result.Ranklustering, type = "Array")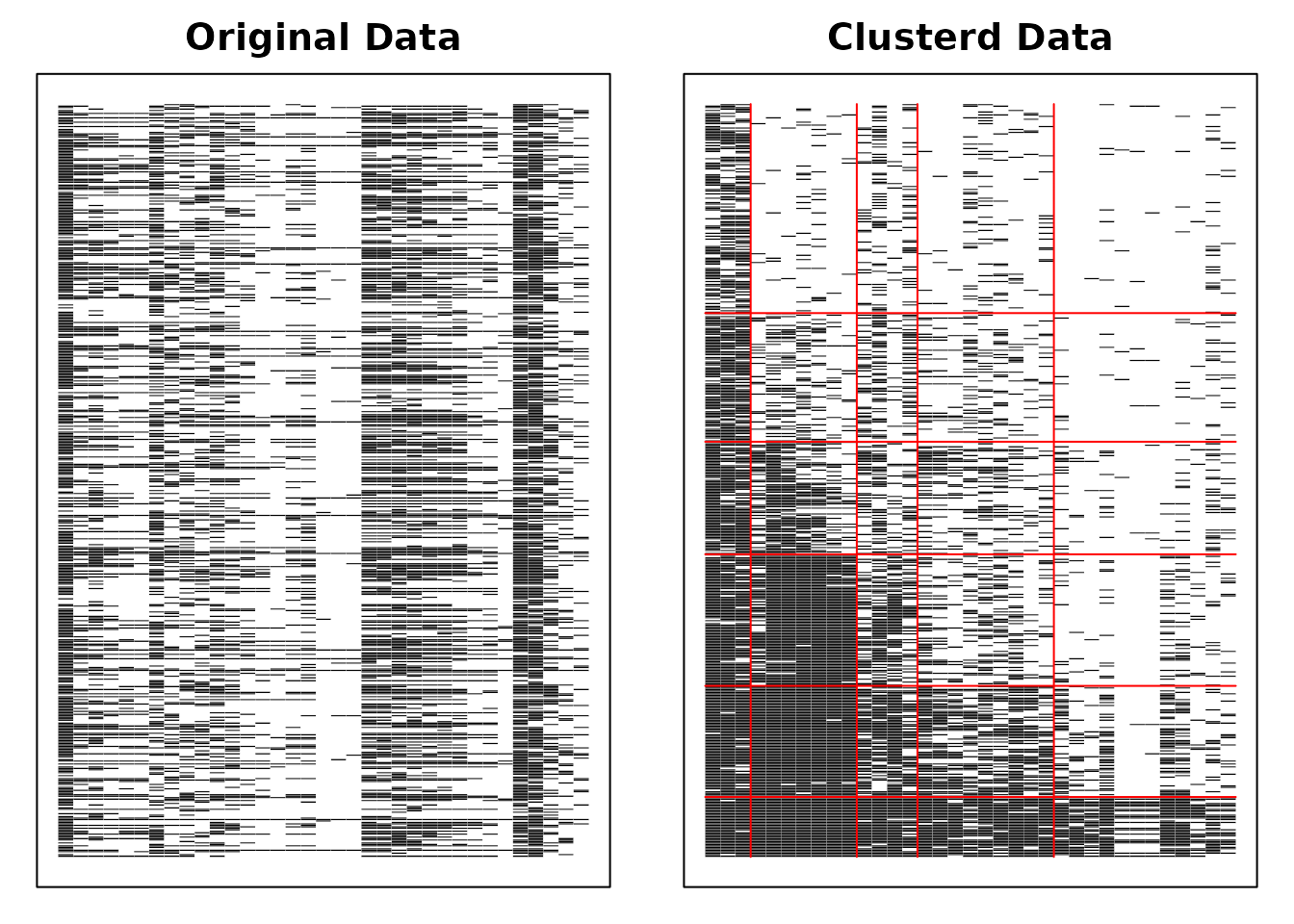
plot(result.Ranklustering, type = "FRP", nc = 2, nr = 3)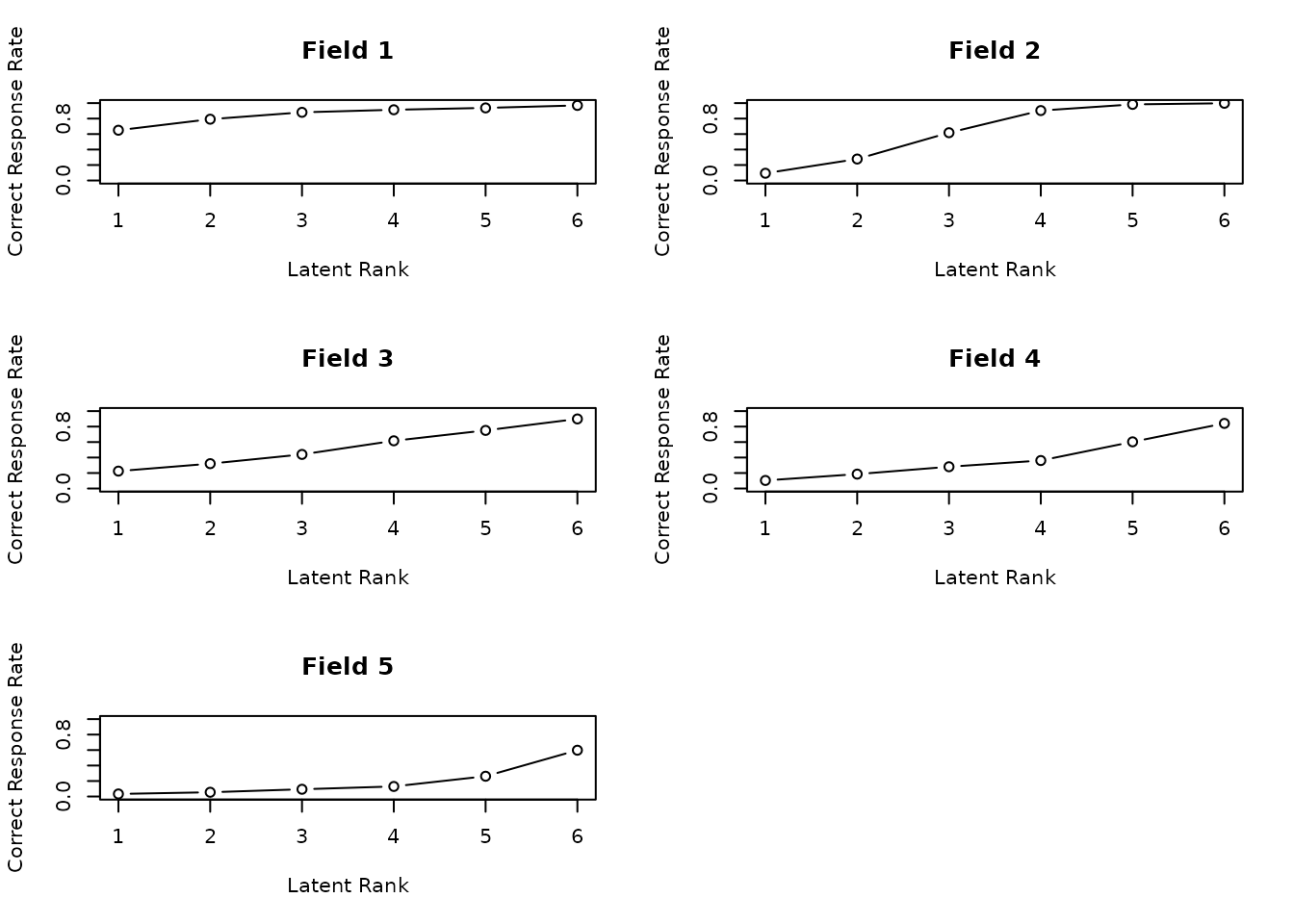
plot(result.Ranklustering, type = "RRV")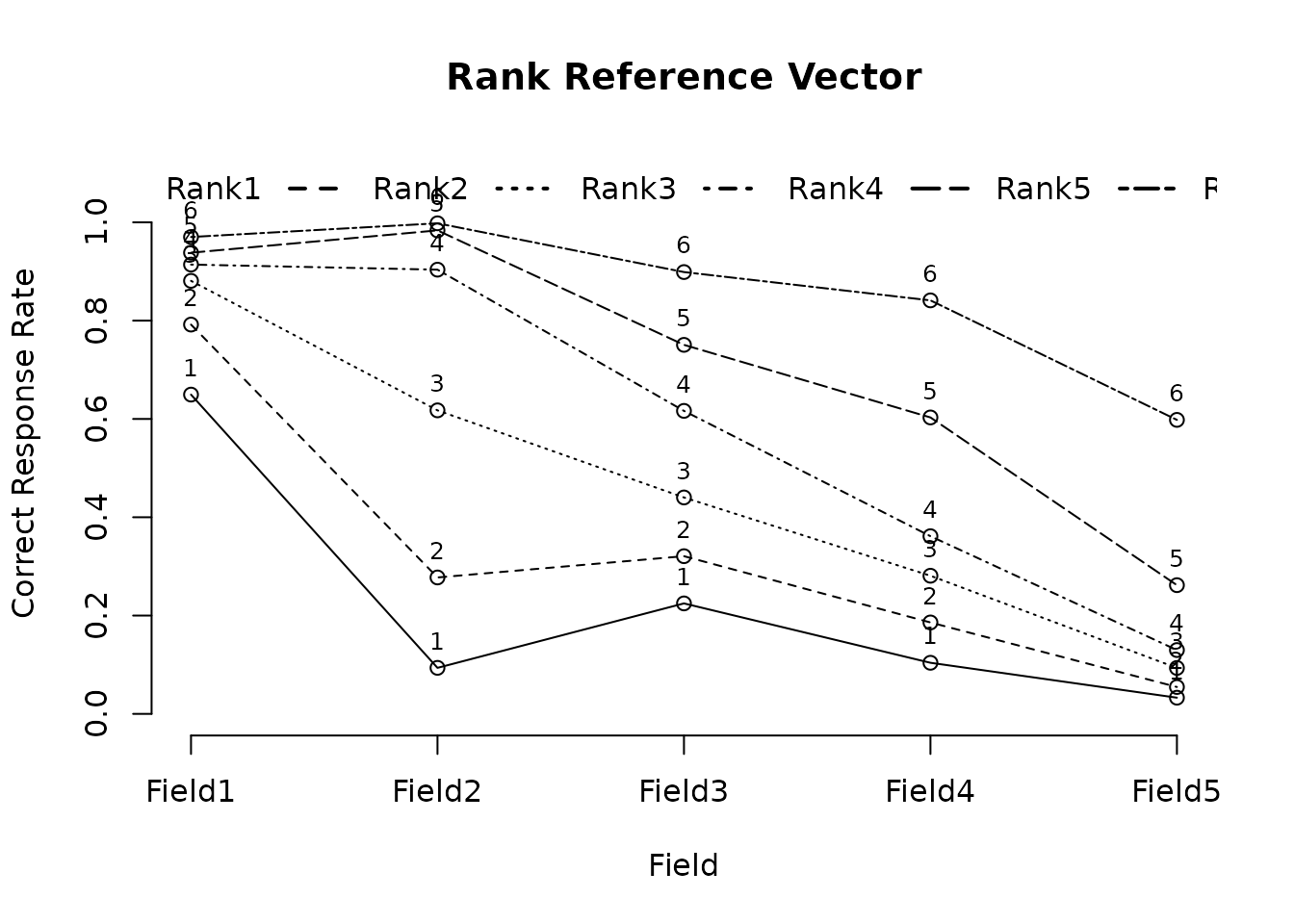
plot(result.Ranklustering, type = "RMP", students = 1:9, nc = 3, nr = 3)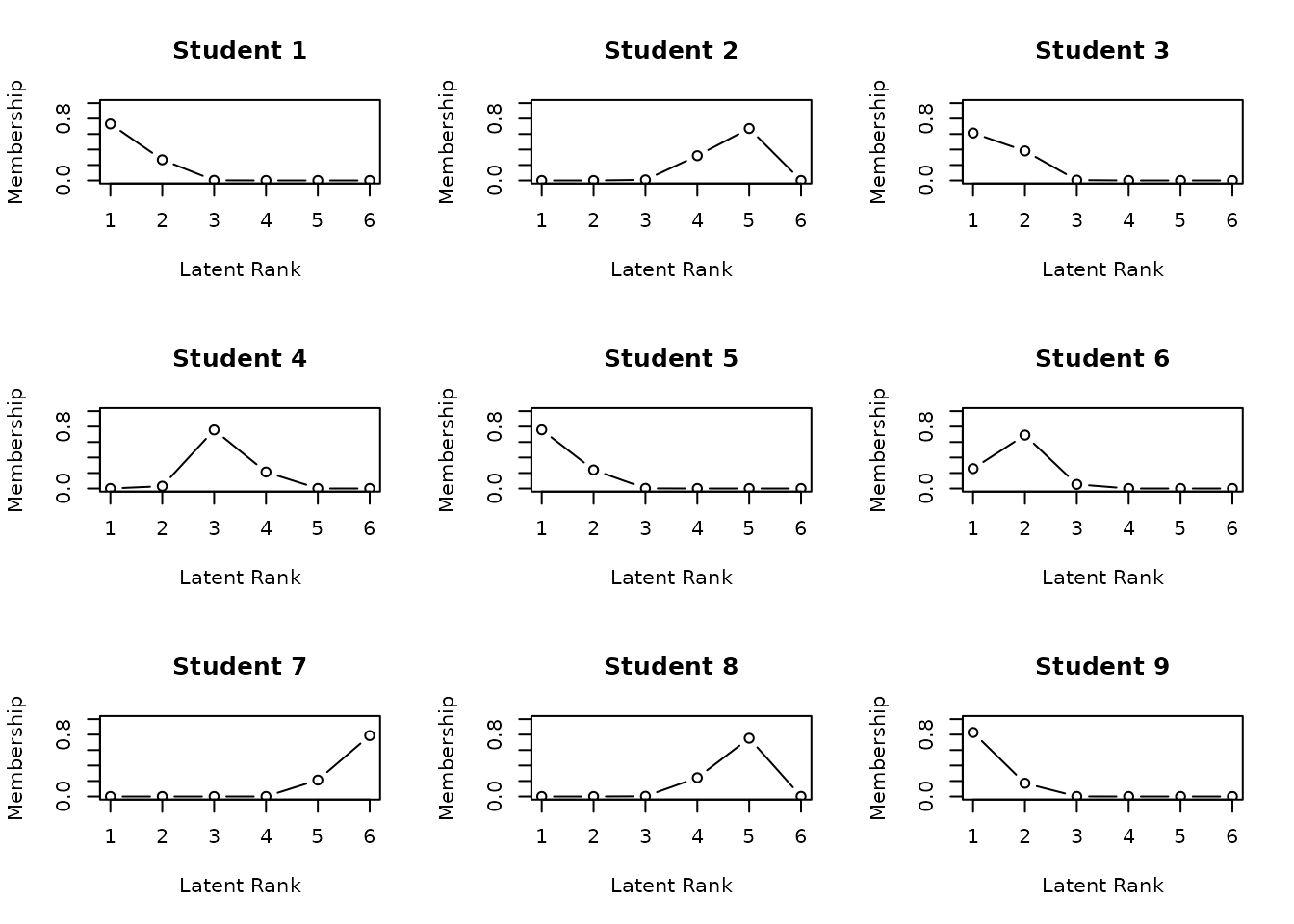
plot(result.Ranklustering, type = "LRD")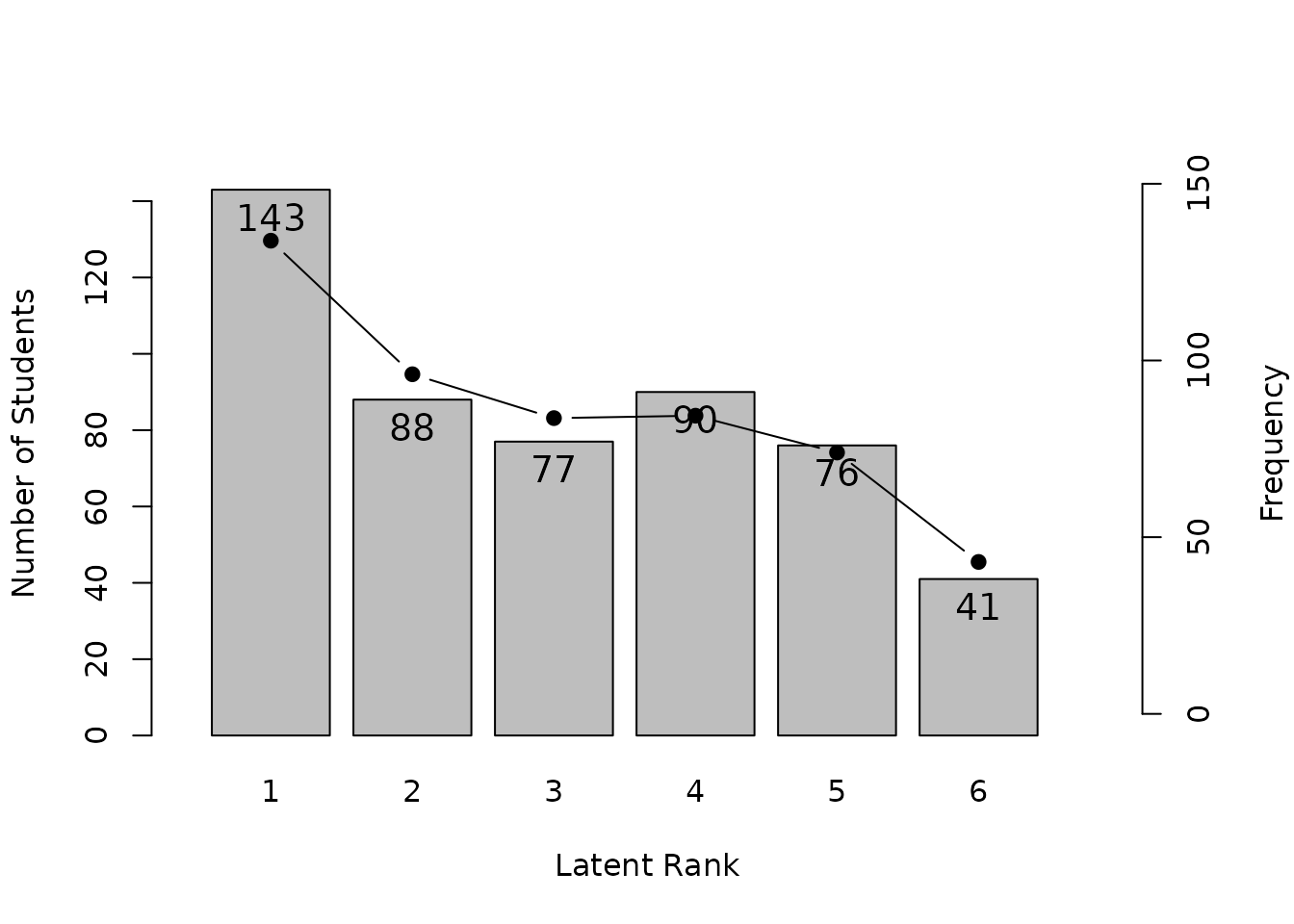
Finding Optimal Number of Classes and Fields
Grid Search
GridSearch() systematically evaluates multiple parameter
combinations and selects the best-fitting model:
result <- GridSearch(J35S515, method = "R", max_ncls = 10, max_nfld = 10, index = "BIC")
result$optimal_ncls
result$optimal_nfld
result$optimal_resultInfinite Relational Model (IRM)
The IRM uses the Chinese Restaurant Process to automatically determine the optimal number of fields and classes:
result.IRM <- Biclustering_IRM(J35S515, gamma_c = 1, gamma_f = 1, verbose = TRUE)
plot(result.IRM, type = "Array")
plot(result.IRM, type = "FRP", nc = 3)
plot(result.IRM, type = "TRP")Biclustering for Polytomous Data
Ordinal Data
result.B.ord <- Biclustering(J35S500, ncls = 5, nfld = 5, method = "R")
result.B.ord
plot(result.B.ord, type = "Array")FRP (Field Reference Profile) shows the expected score per field across latent ranks:
plot(result.B.ord, type = "FRP", nc = 3, nr = 2)FCRP (Field Category Response Profile) shows category probabilities
across ranks. The style parameter can be
"line" or "bar":
plot(result.B.ord, type = "FCRP", nc = 3, nr = 2)
plot(result.B.ord, type = "FCRP", style = "bar", nc = 3, nr = 2)FCBR (Field Cumulative Boundary Reference) shows cumulative boundary probabilities (ordinal only):
plot(result.B.ord, type = "FCBR", nc = 3, nr = 2)ScoreField and RRV plots:
Nominal Data
result.B.nom <- Biclustering(J20S600, ncls = 5, nfld = 4)
result.B.nom
plot(result.B.nom, type = "Array")Nominal Biclustering supports FRP, FCRP, ScoreField, and RRV (but not FCBR):
Rated Data (Multiple-Choice with Correct Answers)
Rated data has both multiple response categories and correct answers.
Biclustering.rated() internally runs nominal Biclustering,
then sorts classes by correct response rate:
result.B.rated <- Biclustering(J21S300, ncls = 5, nfld = 3, method = "R", maxiter = 300)
result.B.rated
#> Ranklustering Analysis (Rated) [MIC]
#>
#> Ranklustering Reference Matrix Profile (Nominal)
#> For category 1
#> Rank 1 Rank 2 Rank 3 Rank 4 Rank 5
#> Field 1 0.106 0.350 0.2827 0.6621 0.7081
#> Field 2 0.257 0.242 0.1836 0.0714 0.0754
#> Field 3 0.267 0.221 0.0967 0.1615 0.0041
#> For category 2
#> Rank 1 Rank 2 Rank 3 Rank 4 Rank 5
#> Field 1 0.283 0.257 0.228 0.1220 0.0974
#> Field 2 0.118 0.291 0.415 0.7994 0.7760
#> Field 3 0.308 0.216 0.129 0.0691 0.0602
#> For category 3
#> Rank 1 Rank 2 Rank 3 Rank 4 Rank 5
#> Field 1 0.314 0.182 0.237 0.1165 0.1030
#> Field 2 0.300 0.257 0.191 0.0665 0.0777
#> Field 3 0.121 0.411 0.555 0.7038 0.8299
#> For category 4
#> Rank 1 Rank 2 Rank 3 Rank 4 Rank 5
#> Field 1 0.297 0.211 0.252 0.0994 0.0915
#> Field 2 0.325 0.210 0.211 0.0627 0.0709
#> Field 3 0.304 0.153 0.219 0.0657 0.1059
#>
#> Field Reference Profile (Binary)
#> Rank 1 Rank 2 Rank 3 Rank 4 Rank 5
#> Field 1 0.102 0.390 0.252 0.660 0.702
#> Field 2 0.115 0.280 0.424 0.803 0.775
#> Field 3 0.121 0.393 0.574 0.657 0.835
#>
#> Field Reference Profile Indices
#> Alpha A Beta B Gamma C
#> Field1 3 0.409 2 0.390 0.25 -0.1383
#> Field2 3 0.379 3 0.424 0.25 -0.0285
#> Field3 1 0.272 3 0.574 0.00 0.0000
#>
#> Rank 1 Rank 2 Rank 3 Rank 4 Rank 5
#> Test Reference Profile 2.370 7.442 8.746 14.844 16.183
#> Latent Rank Ditribution 73.000 52.000 59.000 45.000 71.000
#> Rank Membership Distribution 69.457 57.704 59.047 57.877 55.914
#>
#> Field Membership Profile
#> CRR LFE Field1 Field2 Field3
#> Item01 0.440 1 1.000 0.000 0.000
#> Item02 0.430 1 1.000 0.000 0.000
#> Item07 0.417 1 1.000 0.000 0.000
#> Item06 0.407 1 1.000 0.000 0.000
#> Item03 0.400 1 1.000 0.000 0.000
#> Item04 0.380 1 1.000 0.000 0.000
#> Item05 0.377 1 1.000 0.000 0.000
#> Item09 0.510 2 0.000 1.000 0.000
#> Item13 0.473 2 0.000 1.000 0.000
#> Item12 0.470 2 0.000 1.000 0.000
#> Item14 0.463 2 0.000 1.000 0.000
#> Item11 0.460 2 0.000 1.000 0.000
#> Item08 0.450 2 0.000 1.000 0.000
#> Item10 0.420 2 0.000 1.000 0.000
#> Item18 0.530 3 0.000 0.000 1.000
#> Item21 0.527 3 0.000 0.000 1.000
#> Item16 0.520 3 0.000 0.000 1.000
#> Item15 0.510 3 0.000 0.000 1.000
#> Item19 0.500 3 0.000 0.000 1.000
#> Item20 0.497 3 0.000 0.000 1.000
#> Item17 0.463 3 0.000 0.000 1.000
#> Latent Field Distribution
#> Field 1 Field 2 Field 3
#> N of Items 7 7 7
#>
#> Model Fit Indices (Binary)
#> Number of Latent Rank : 5
#> Number of Latent Field: 3
#> Number of EM cycle: 296
#> value
#> model_log_like -3293.331
#> bench_log_like -3079.142
#> null_log_like -4318.046
#> model_Chi_sq 428.376
#> null_Chi_sq 2477.806
#> model_df 336.000
#> null_df 420.000
#> NFI 0.827
#> RFI 0.784
#> IFI 0.957
#> TLI 0.944
#> CFI 0.955
#> RMSEA 0.030
#> AIC -243.624
#> CAIC -1824.094
#> BIC -1488.094
#>
#> Model Fit Indices (Nominal)
#> value
#> model_log_like -7013.548
#> AIC 14117.095
#> CAIC 14328.766
#> BIC 14283.766
#> Weakly Ordinal Alignment Condition is Satisfied.
plot(result.B.rated, type = "Array")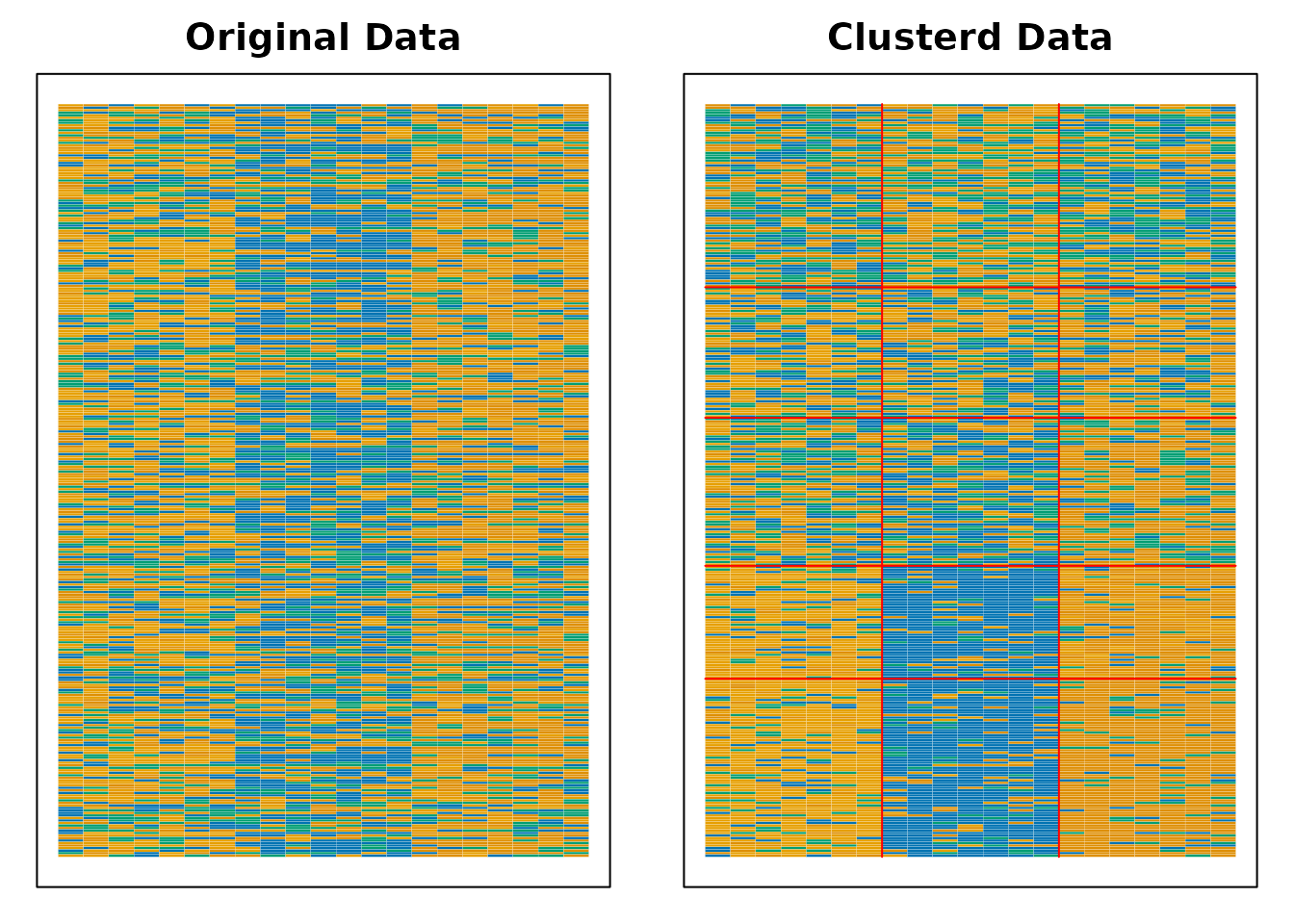
plot(result.B.rated, type = "FRP", nc = 3, nr = 1)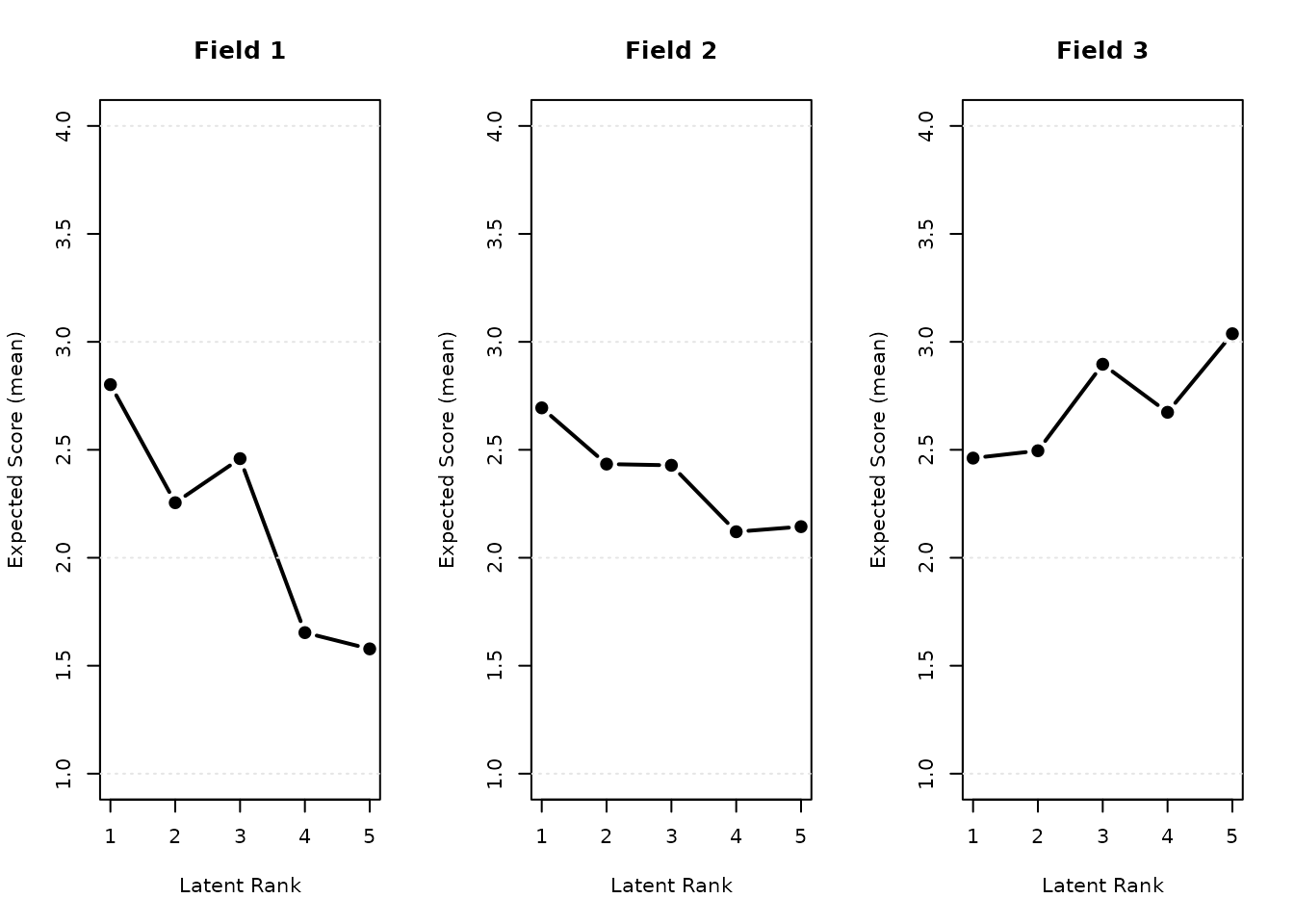
Two layers of fit indices are reported: binary (with CFI/RMSEA) and
nominal (AIC/BIC/CAIC only). Access them via
result$TestFitIndices and
result$TestFitIndices_nominal.
Rated IRM
The IRM also supports rated data:
result.IRM.rated <- Biclustering_IRM(J21S300, gamma_c = 1, gamma_f = 1, verbose = FALSE)
plot(result.IRM.rated, type = "Array")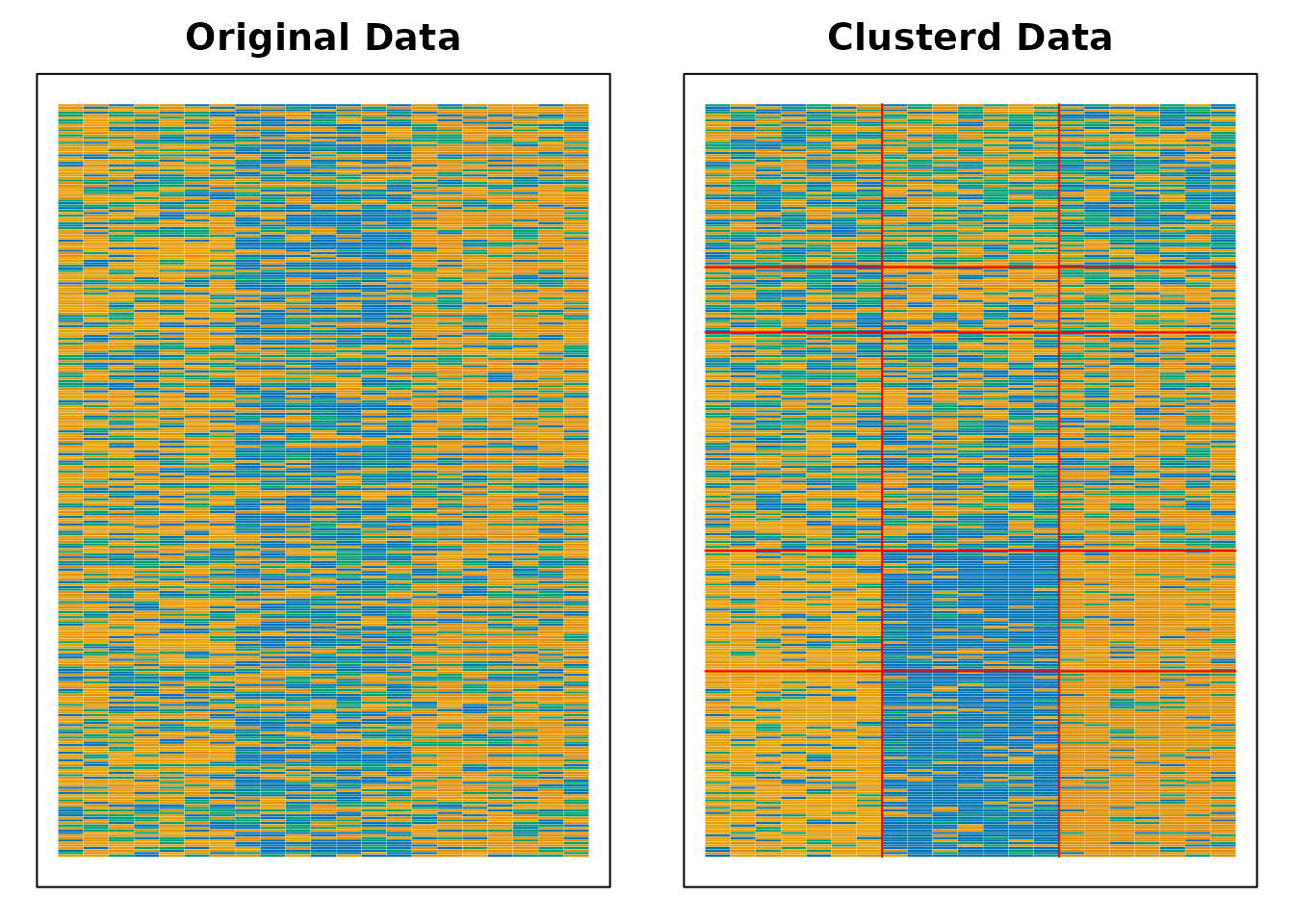
plot(result.IRM.rated, type = "FRP", nc = 3, nr = 1)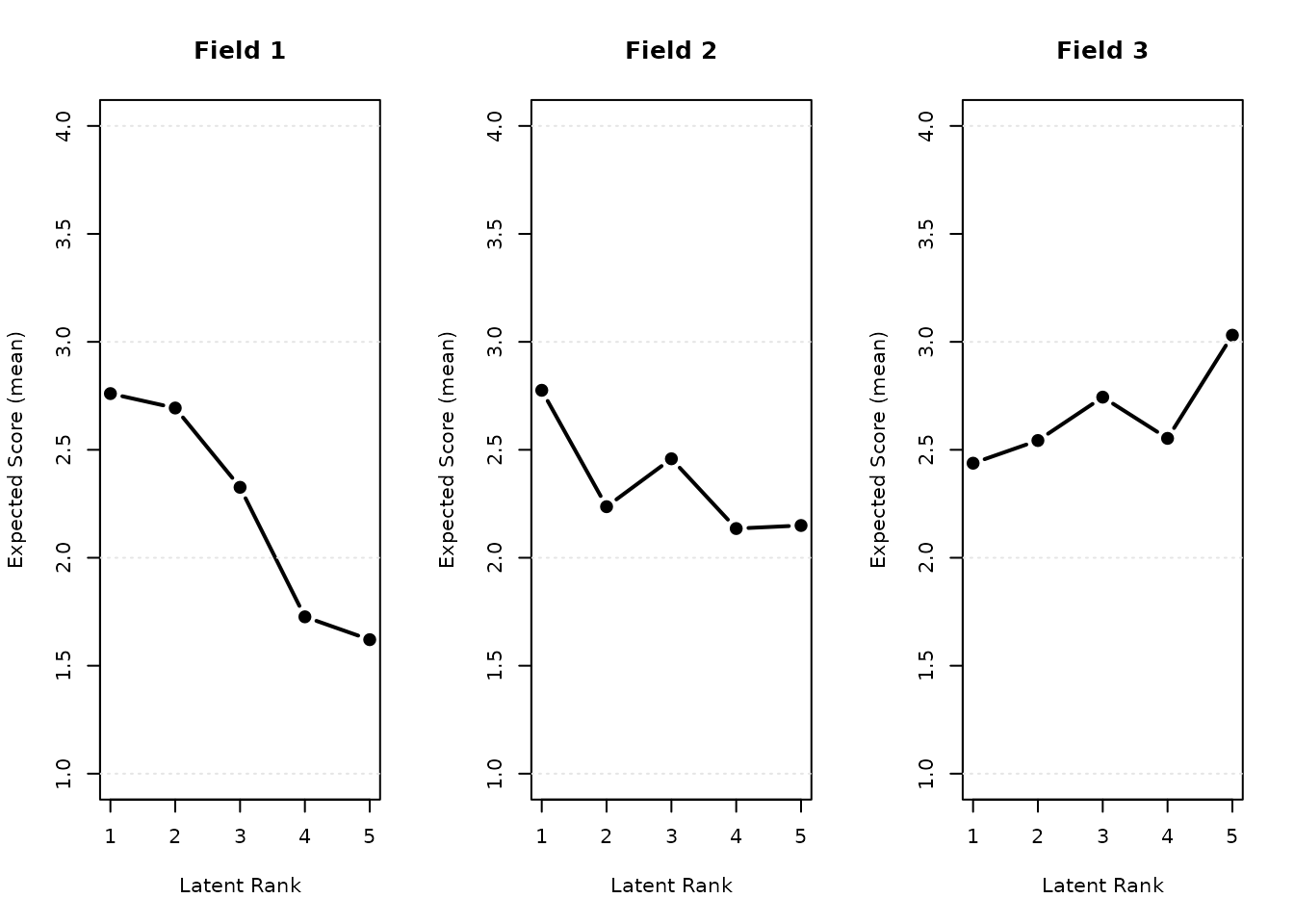
Distractor Analysis
DistractorAnalysis() examines how examinees in each
rank/class respond to each item’s categories. It computes observed
frequencies, proportions, chi-square statistics, p-values, and Cramer’s
V (effect size) for each item-by-rank cell.
result.B.rated <- Biclustering(J21S300, ncls = 5, nfld = 3, method = "R", maxiter = 300)
da <- DistractorAnalysis(result.B.rated)
# Full output (grouped by field for Biclustering)
print(da)
# Filter by items and/or ranks
print(da, items = 1:7, ranks = c(1, 5))
# Plot distractor bar charts
plot(da, items = 1:6, nc = 3, nr = 2)DistractorAnalysis() also works with LRA()
results for rated data:
result.LRA.rated <- LRA(J21S300, nrank = 5, mic = TRUE)
da_lra <- DistractorAnalysis(result.LRA.rated)
print(da_lra, items = 1:3)
plot(da_lra, items = 1:6, nc = 3, nr = 2)