
Bayesian Network and Local Dependence Models
Source:vignettes/network-models.Rmd
network-models.RmdNote: Most examples in this vignette are shown with
eval=FALSEto keep CRAN build times short. For full rendered output, see the pkgdown site.
Bayesian Network Model (BNM)
BNM represents conditional probabilities between items in a network
format. A Directed Acyclic Graph (DAG) must be provided via
adj_matrix, adj_file, or g
(igraph object).
Creating the Graph
DAG <- matrix(
c(
"Item01", "Item02",
"Item02", "Item03",
"Item02", "Item04",
"Item03", "Item05",
"Item04", "Item05"
),
ncol = 2, byrow = TRUE
)
# Graph object
g <- igraph::graph_from_data_frame(DAG)
g
#> IGRAPH d443084 DN-- 5 5 --
#> + attr: name (v/c)
#> + edges from d443084 (vertex names):
#> [1] Item01->Item02 Item02->Item03 Item02->Item04 Item03->Item05 Item04->Item05
# Adjacency matrix
adj_mat <- as.matrix(igraph::get.adjacency(g))
print(adj_mat)
#> Item01 Item02 Item03 Item04 Item05
#> Item01 0 1 0 0 0
#> Item02 0 0 1 1 0
#> Item03 0 0 0 0 1
#> Item04 0 0 0 0 1
#> Item05 0 0 0 0 0Running BNM
result.BNM <- BNM(J5S10, adj_matrix = adj_mat)
result.BNM
#> Adjacency Matrix
#> Item01 Item02 Item03 Item04 Item05
#> Item01 0 1 0 0 0
#> Item02 0 0 1 1 0
#> Item03 0 0 0 0 1
#> Item04 0 0 0 0 1
#> Item05 0 0 0 0 0
#> [1] "Your graph is an acyclic graph."
#> [1] "Your graph is connected DAG."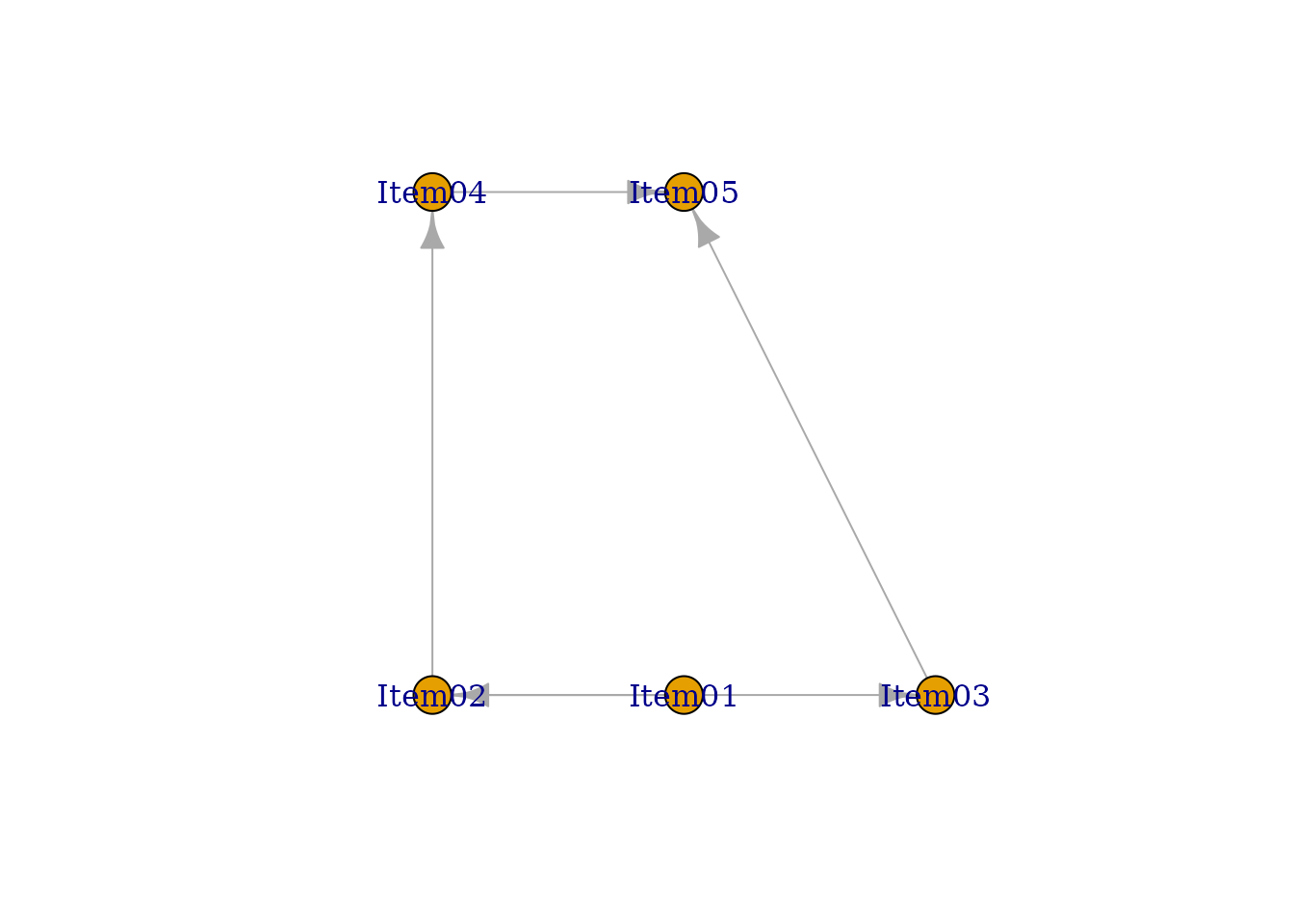
#>
#> Parameter Learning
#> PIRP 1 PIRP 2 PIRP 3 PIRP 4
#> Item01 0.600
#> Item02 0.250 0.5
#> Item03 0.833 1.0
#> Item04 0.167 0.5
#> Item05 0.000 NaN 0.333 0.667
#>
#> Conditional Correct Response Rate
#> Child Item N of Parents Parent Items PIRP Conditional CRR
#> 1 Item01 0 No Parents No Pattern 0.6000000
#> 2 Item02 1 Item01 0 0.2500000
#> 3 Item02 1 Item01 1 0.5000000
#> 4 Item03 1 Item02 0 0.8333333
#> 5 Item03 1 Item02 1 1.0000000
#> 6 Item04 1 Item02 0 0.1666667
#> 7 Item04 1 Item02 1 0.5000000
#> 8 Item05 2 Item03, Item04 00 0.0000000
#> 9 Item05 2 Item03, Item04 01 NaN(0/0)
#> 10 Item05 2 Item03, Item04 10 0.3333333
#> 11 Item05 2 Item03, Item04 11 0.6666667
#>
#> Model Fit Indices
#> value
#> model_log_like -27.046
#> bench_log_like -8.935
#> null_log_like -28.882
#> model_Chi_sq 36.222
#> null_Chi_sq 39.894
#> model_df 20.000
#> null_df 25.000
#> NFI 0.092
#> RFI 0.000
#> IFI 0.185
#> TLI 0.000
#> CFI 0.000
#> RMSEA 0.300
#> AIC -3.778
#> CAIC -29.829
#> BIC -9.829Structure Learning with Genetic Algorithm
BNM_GA() searches for a DAG suitable for the data using
a genetic algorithm:
BNM_GA(J5S10,
population = 20, Rs = 0.5, Rm = 0.002, maxParents = 2,
maxGeneration = 100, crossover = 2, elitism = 2
)Structure Learning with PBIL
BNM_PBIL(J5S10,
population = 20, Rs = 0.5, Rm = 0.005, maxParents = 2,
alpha = 0.05, estimate = 4
)Local Dependence Latent Rank Analysis (LDLRA)
LDLRA combines LRA and BNM to analyze how item dependency networks change across latent ranks. A graph must be specified for each rank.
Setting Up Rank-Specific Graphs
Graphs can be provided via CSV file, adjacency matrix list, or igraph object list:
DAG_dat <- matrix(c(
"From", "To", "Rank",
"Item01", "Item02", 1,
"Item04", "Item05", 1,
"Item01", "Item02", 2,
"Item02", "Item03", 2,
"Item04", "Item05", 2,
"Item08", "Item09", 2,
"Item08", "Item10", 2,
"Item09", "Item10", 2,
"Item08", "Item11", 2,
"Item01", "Item02", 3,
"Item02", "Item03", 3,
"Item04", "Item05", 3,
"Item08", "Item09", 3,
"Item08", "Item10", 3,
"Item09", "Item10", 3,
"Item08", "Item11", 3,
"Item02", "Item03", 4,
"Item04", "Item06", 4,
"Item04", "Item07", 4,
"Item05", "Item06", 4,
"Item05", "Item07", 4,
"Item08", "Item10", 4,
"Item08", "Item11", 4,
"Item09", "Item11", 4,
"Item02", "Item03", 5,
"Item04", "Item06", 5,
"Item04", "Item07", 5,
"Item05", "Item06", 5,
"Item05", "Item07", 5,
"Item09", "Item11", 5,
"Item10", "Item11", 5,
"Item10", "Item12", 5
), ncol = 3, byrow = TRUE)
edgeFile <- tempfile(fileext = ".csv")
write.csv(DAG_dat, edgeFile, row.names = FALSE, quote = TRUE)Running LDLRA
result.LDLRA <- LDLRA(J12S5000, ncls = 5, adj_file = edgeFile)
result.LDLRAStructure Learning for LDLRA with PBIL
LDLRA_PBIL() learns item-interaction graphs for each
rank automatically:
result.LDLRA.PBIL <- LDLRA_PBIL(J35S515,
seed = 123, ncls = 5, method = "R",
elitism = 1, successiveLimit = 15
)
result.LDLRA.PBILLocal Dependence Biclustering (LDB)
LDB combines biclustering with Bayesian network models, analyzing relationships between item fields within each rank.
conf <- c(
1, 6, 6, 8, 9, 9, 4, 7, 7, 7, 5, 8, 9, 10, 10,
9, 9, 10, 10, 10, 2, 2, 3, 3, 5, 5, 6, 9, 9, 10,
1, 1, 7, 9, 10
)
edges_data <- data.frame(
"From Field (Parent) >>>" = c(
6, 4, 5, 1, 1, 4,
3, 4, 6, 2, 4, 4,
3, 6, 4, 1,
7, 9, 6, 7
),
">>> To Field (Child)" = c(
8, 7, 8, 7, 2, 5,
5, 8, 8, 4, 6, 7,
5, 8, 5, 8,
10, 10, 8, 9
),
"At Class/Rank (Locus)" = c(
2, 2, 2, 2, 2, 2,
3, 3, 3, 3, 3, 3,
4, 4, 4, 4,
5, 5, 5, 5
)
)
edgeFile <- tempfile(fileext = ".csv")
write.csv(edges_data, file = edgeFile, row.names = FALSE)
result.LDB <- LDB(U = J35S515, ncls = 5, conf = conf, adj_file = edgeFile)
result.LDB
plot(result.LDB, type = "Array")
plot(result.LDB, type = "TRP")
plot(result.LDB, type = "LRD")
plot(result.LDB, type = "RMP", students = 1:9, nc = 3, nr = 3)
plot(result.LDB, type = "FRP", nc = 3, nr = 2)FieldPIRP visualizes correct answer counts for each rank and field:
plot(result.LDB, type = "FieldPIRP")Bicluster Network Model (BINET)
BINET combines biclustering with class-level network analysis. Unlike LDB where nodes are fields, in BINET the nodes represent classes.
conf <- c(
1, 5, 5, 5, 9, 9, 6, 6, 6, 6, 2, 7, 7, 11, 11,
7, 7, 12, 12, 12, 2, 2, 3, 3, 4, 4, 4, 8, 8, 12,
1, 1, 6, 10, 10
)
edges_data <- data.frame(
"From Class (Parent) >>>" = c(
1, 2, 3, 4, 5, 7,
2, 4, 6, 8, 10,
6, 6, 11, 8, 9, 12
),
">>> To Class (Child)" = c(
2, 4, 5, 5, 6, 11,
3, 7, 9, 12, 12,
10, 8, 12, 12, 11, 13
),
"At Field (Locus)" = c(
1, 2, 2, 3, 4, 4,
5, 5, 5, 5, 5,
7, 8, 8, 9, 9, 12
)
)
edgeFile <- tempfile(fileext = ".csv")
write.csv(edges_data, file = edgeFile, row.names = FALSE)
result.BINET <- BINET(
U = J35S515, ncls = 13, nfld = 12,
conf = conf, adj_file = edgeFile
)
print(result.BINET)
plot(result.BINET, type = "Array")
plot(result.BINET, type = "TRP")
plot(result.BINET, type = "LRD")
plot(result.BINET, type = "RMP", students = 1:9, nc = 3, nr = 3)
plot(result.BINET, type = "FRP", nc = 3, nr = 2)LDPSR shows Passing Student Rates for locally dependent classes:
plot(result.BINET, type = "LDPSR", nc = 3, nr = 2)