This function takes exametrika output as input and generates array plots using ggplot2. Array plots visualize the response patterns of students (rows) across items (columns), showing both the original data and the clustered/reordered data.
Supports both binary (0/1) and multi-valued (ordinal/nominal) data.
Usage
plotArray_gg(
data,
Original = TRUE,
Clusterd = TRUE,
Clusterd_lines = TRUE,
Clusterd_lines_color = NULL,
title = TRUE,
colors = NULL,
show_legend = NULL,
legend_position = "right"
)Arguments
- data
An object of class
c("exametrika", "Biclustering"),c("exametrika", "nominalBiclustering"),c("exametrika", "ordinalBiclustering"),c("exametrika", "ratedBiclustering"),c("exametrika", "IRM"),c("exametrika", "LDB"), orc("exametrika", "BINET").- Original
Logical. If
TRUE(default), plot the original (unsorted) response data.- Clusterd
Logical. If
TRUE(default), plot the clustered (sorted by class and field) response data.- Clusterd_lines
Logical. If
TRUE(default), draw lines on the clustered plot to indicate class and field boundaries.- Clusterd_lines_color
Character. Color of the boundary lines. If
NULL(default), uses"red"for binary data or"white"for multi-valued data.- title
Logical or character. If
TRUE(default), display auto-generated titles. IfFALSE, no titles. If a character string, use it as a custom title prefix.- colors
Character vector of colors for each category. If
NULL(default), uses white/black for binary data or a colorblind-friendly palette for multi-valued data.- show_legend
Logical. If
TRUE, display the legend. Default isFALSEfor binary data,TRUEfor multi-valued.- legend_position
Character. Position of the legend. One of
"right"(default),"top","bottom","left","none".
Value
A ggplot object or a grid arrangement of two ggplot objects (original and clustered plots side by side).
Details
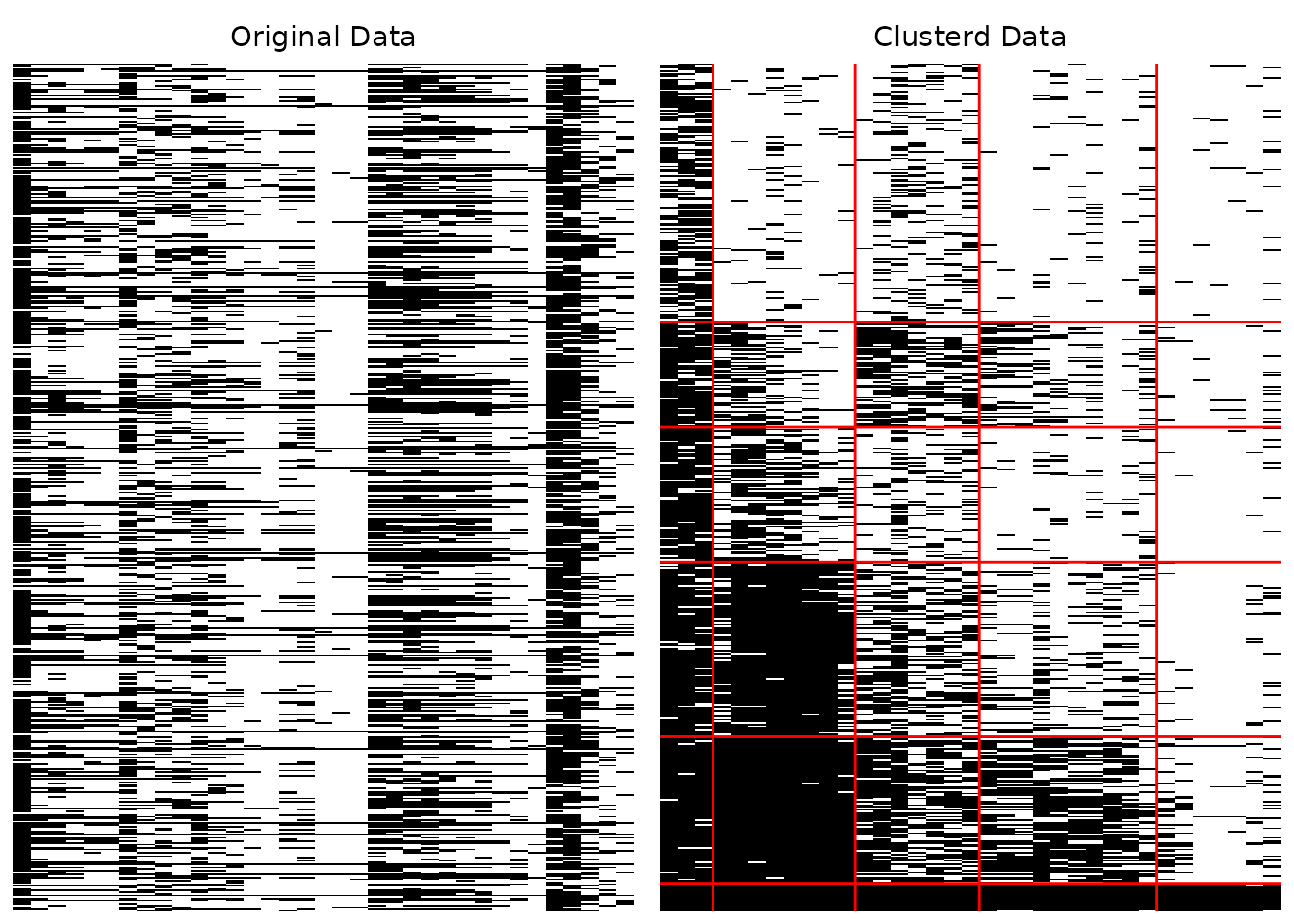
The array plot provides a visual representation of the biclustering result. For binary data, black cells indicate correct responses (1) and white cells indicate incorrect responses (0). For multi-valued data, different colors represent different response categories.
In a well-fitted model, the clustered plot should show a clear block diagonal pattern where high-ability students (bottom rows) answer difficult items (right columns) correctly.
Examples
library(exametrika)
result <- Biclustering(J35S515, nfld = 5, ncls = 6)
#> Biclustering is chosen.
#>
iter 1 log_lik -7966.66
#>
iter 2 log_lik -7442.38
#>
iter 3 log_lik -7266.35
#>
iter 4 log_lik -7151.01
#>
iter 5 log_lik -7023.94
#>
iter 6 log_lik -6984.82
#>
iter 7 log_lik -6950.27
#>
iter 8 log_lik -6939.34
#>
iter 9 log_lik -6930.89
#>
iter 10 log_lik -6923.5
#>
iter 11 log_lik -6914.56
#>
iter 12 log_lik -6908.89
#>
iter 13 log_lik -6906.84
#>
iter 14 log_lik -6905.39
#>
iter 15 log_lik -6904.24
#>
iter 16 log_lik -6903.28
#>
iter 17 log_lik -6902.41
#>
iter 18 log_lik -6901.58
#>
iter 19 log_lik -6900.74
#>
iter 20 log_lik -6899.86
#>
iter 21 log_lik -6898.9
#>
iter 22 log_lik -6897.84
#>
iter 23 log_lik -6896.66
#>
iter 24 log_lik -6895.35
#>
iter 25 log_lik -6893.92
#>
iter 26 log_lik -6892.4
#>
iter 27 log_lik -6890.85
#>
iter 28 log_lik -6889.32
#>
iter 29 log_lik -6887.9
#>
iter 30 log_lik -6886.66
#>
iter 31 log_lik -6885.67
#>
iter 32 log_lik -6884.98
#>
iter 33 log_lik -6884.58
#>
#> No ID column detected. All columns treated as response data. Sequential IDs (Student1, Student2, ...) were generated. Use id= parameter to specify the ID column explicitly.
# Basic usage
plotArray_gg(result)
 #> TableGrob (1 x 2) "arrange": 2 grobs
#> z cells name grob
#> 1 1 (1-1,1-1) arrange gtable[layout]
#> 2 2 (1-1,2-2) arrange gtable[layout]
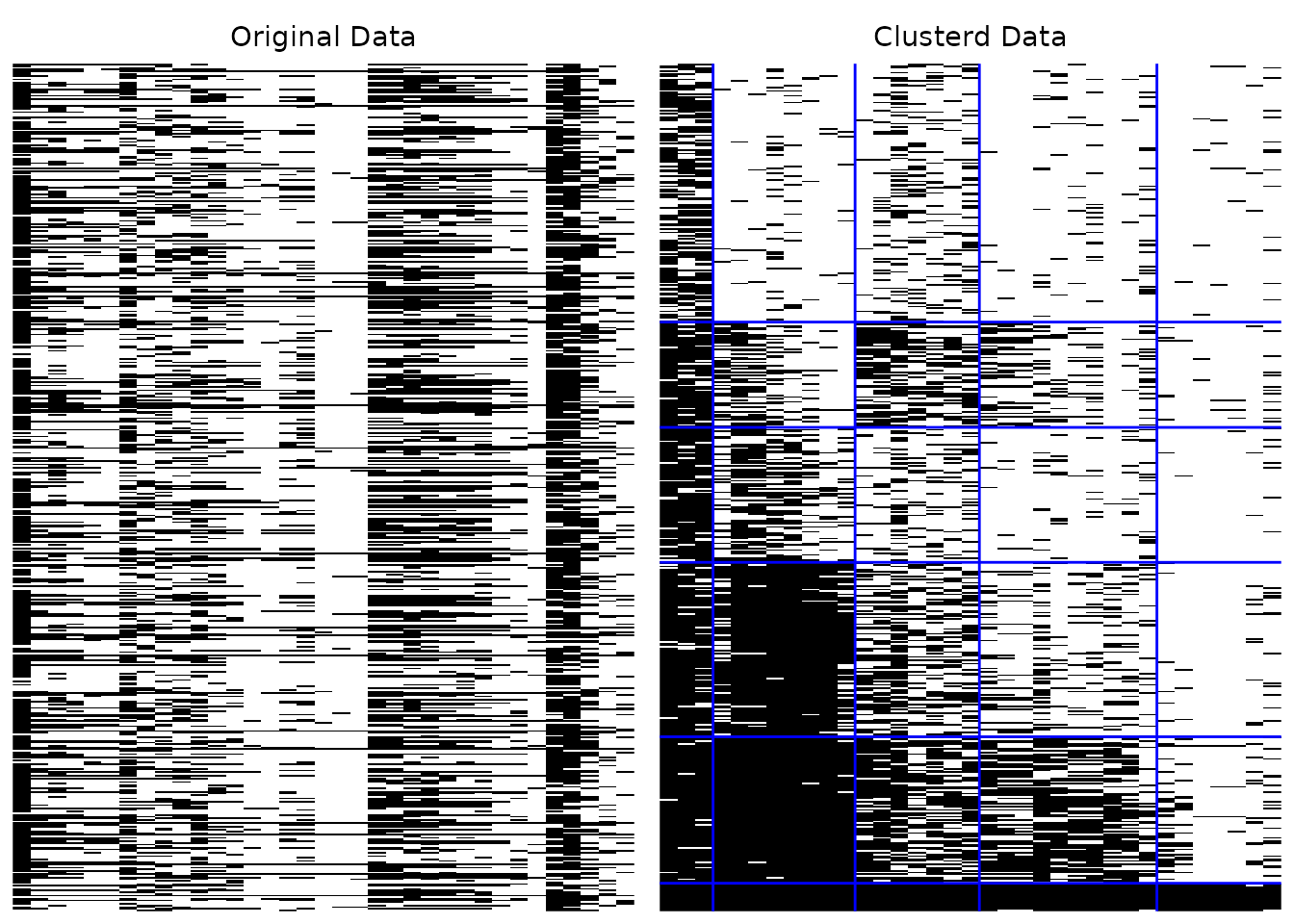
# Custom boundary line color
plotArray_gg(result, Clusterd_lines_color = "blue")
#> TableGrob (1 x 2) "arrange": 2 grobs
#> z cells name grob
#> 1 1 (1-1,1-1) arrange gtable[layout]
#> 2 2 (1-1,2-2) arrange gtable[layout]
# Custom boundary line color
plotArray_gg(result, Clusterd_lines_color = "blue")
 #> TableGrob (1 x 2) "arrange": 2 grobs
#> z cells name grob
#> 1 1 (1-1,1-1) arrange gtable[layout]
#> 2 2 (1-1,2-2) arrange gtable[layout]
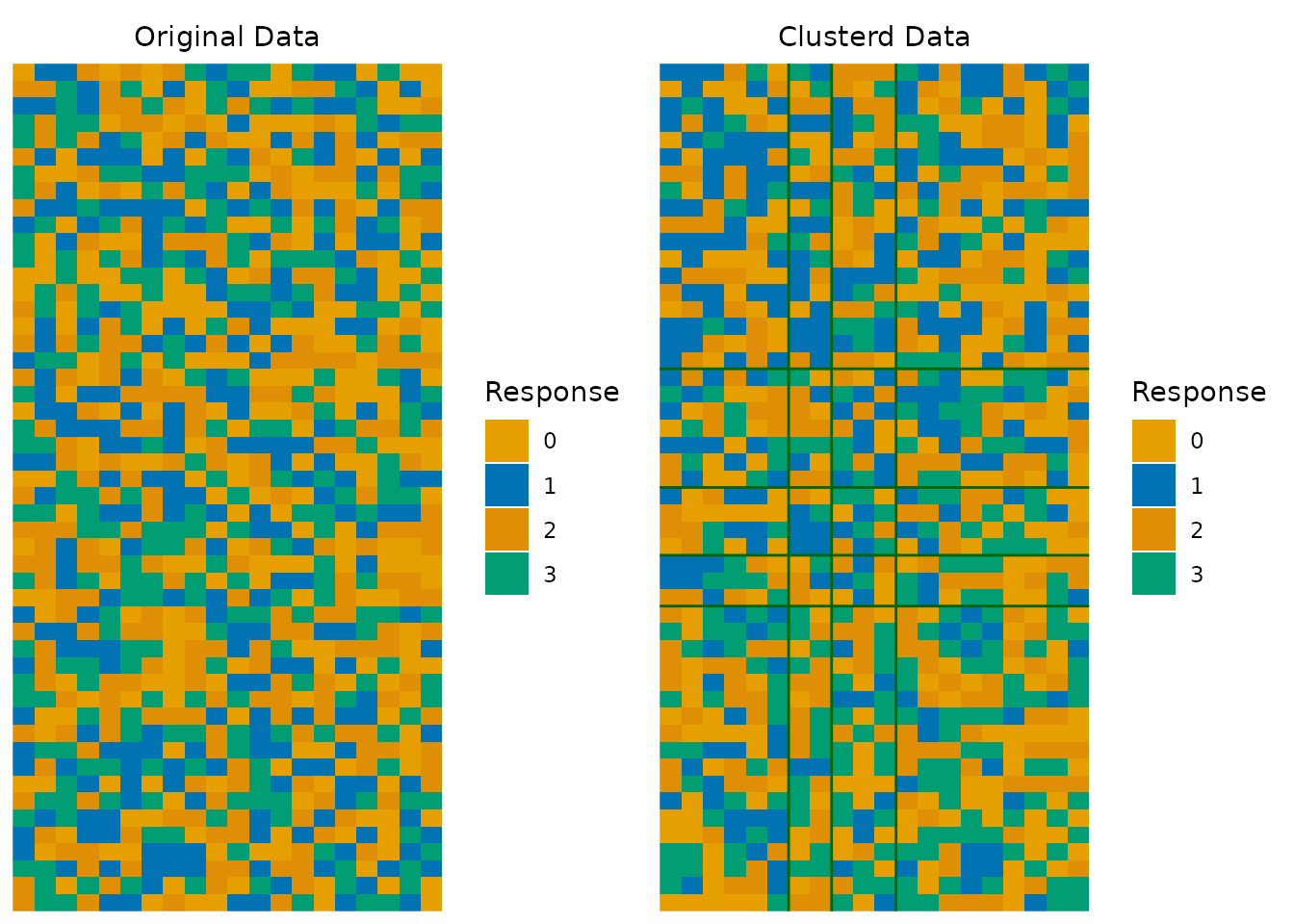
# Multi-valued data with custom colors
synthetic_data <- matrix(sample(0:3, 50 * 20, replace = TRUE), nrow = 50, ncol = 20)
result_multi <- Biclustering(synthetic_data, nfld = 4, ncls = 5)
#> No ID column detected. All columns treated as response data. Sequential IDs (Student1, Student2, ...) were generated. Use id= parameter to specify the ID column explicitly.
#> Biclustering is chosen.
#>
iter 1 log_lik -805.847
#>
iter 2 log_lik -805.895
#> Strongly ordinal alignment condition was satisfied.
plotArray_gg(result_multi, show_legend = TRUE, Clusterd_lines_color = "darkgreen")
#> TableGrob (1 x 2) "arrange": 2 grobs
#> z cells name grob
#> 1 1 (1-1,1-1) arrange gtable[layout]
#> 2 2 (1-1,2-2) arrange gtable[layout]
# Multi-valued data with custom colors
synthetic_data <- matrix(sample(0:3, 50 * 20, replace = TRUE), nrow = 50, ncol = 20)
result_multi <- Biclustering(synthetic_data, nfld = 4, ncls = 5)
#> No ID column detected. All columns treated as response data. Sequential IDs (Student1, Student2, ...) were generated. Use id= parameter to specify the ID column explicitly.
#> Biclustering is chosen.
#>
iter 1 log_lik -805.847
#>
iter 2 log_lik -805.895
#> Strongly ordinal alignment condition was satisfied.
plotArray_gg(result_multi, show_legend = TRUE, Clusterd_lines_color = "darkgreen")
 #> TableGrob (1 x 2) "arrange": 2 grobs
#> z cells name grob
#> 1 1 (1-1,1-1) arrange gtable[layout]
#> 2 2 (1-1,2-2) arrange gtable[layout]
#> TableGrob (1 x 2) "arrange": 2 grobs
#> z cells name grob
#> 1 1 (1-1,1-1) arrange gtable[layout]
#> 2 2 (1-1,2-2) arrange gtable[layout]