This function takes exametrika Biclustering output as input and generates a Rank Reference Vector (RRV) plot using ggplot2. RRV shows how each latent rank performs across fields, with one line per rank.
Supports both binary (2-valued) and polytomous (multi-valued) biclustering models.
For polytomous data, the stat parameter controls how expected scores
are calculated from category probabilities.
Usage
plotRRV_gg(
data,
title = TRUE,
colors = NULL,
linetype = "solid",
show_legend = TRUE,
legend_position = "right",
stat = "mean",
show_labels = NULL
)Arguments
- data
An object of class
c("exametrika", "Biclustering")fromexametrika::Biclustering().- title
Logical or character. If
TRUE(default), display an auto-generated title. IfFALSE, no title. If a character string, use it as a custom title.- colors
Character vector of colors for each rank. If
NULL(default), a colorblind-friendly palette is used.- linetype
Character or numeric vector specifying the line types. If a single value, all lines use that type. If a vector, each rank uses the corresponding type. Default is
"solid".- show_legend
Logical. If
TRUE(default), display the legend.- legend_position
Character. Position of the legend. One of
"right"(default),"top","bottom","left","none".- stat
Character. Statistic for polytomous data:
"mean"(default),"median", or"mode". For binary data, this parameter is ignored."mean": Expected score (sum of category x probability)"median": Median category (cumulative probability >= 0.5)"mode": Most probable category
- show_labels
Logical. If
TRUE, displays rank labels on each point usingggrepelto avoid overlaps. Defaults toFALSEsince the legend already provides rank information.
Details
The Rank Reference Vector is the transpose of the Field Reference Profile (FRP). While FRP shows one plot per field, RRV displays all ranks in a single plot with fields on the x-axis. Each line represents a latent rank, showing its performance pattern across fields.
Binary Data (2 categories):
Y-axis shows "Correct Response Rate" (0.0 to 1.0)
Values represent the probability of correct response
Polytomous Data (3+ categories):
Y-axis shows "Expected Score" (1 to max category)
Values are calculated using the
statparameterHigher scores indicate better performance
RRV is used when latent ranks are ordinal (ordered). For unordered
latent classes, use plotCRV_gg instead.
Examples
# Binary biclustering
library(exametrika)
result <- Biclustering(J15S500, nfld = 3, ncls = 5)
#> Biclustering is chosen.
#>
iter 1 log_lik -4020.83
#>
iter 2 log_lik -3997.76
#>
iter 3 log_lik -3992.39
#>
iter 4 log_lik -3986.8
#>
iter 5 log_lik -3980
#>
iter 6 log_lik -3973.35
#>
iter 7 log_lik -3967.73
#>
iter 8 log_lik -3963.4
#>
iter 9 log_lik -3960.25
#>
iter 10 log_lik -3958.04
#>
iter 11 log_lik -3956.52
#>
iter 12 log_lik -3955.47
#>
iter 13 log_lik -3954.72
#>
iter 14 log_lik -3954.17
#>
iter 15 log_lik -3953.75
#>
iter 16 log_lik -3953.39
#>
#>
#> Weakly ordinal alignment condition was satisfied.
#> No ID column detected. All columns treated as response data. Sequential IDs (Student1, Student2, ...) were generated. Use id= parameter to specify the ID column explicitly.
plotRRV_gg(result)
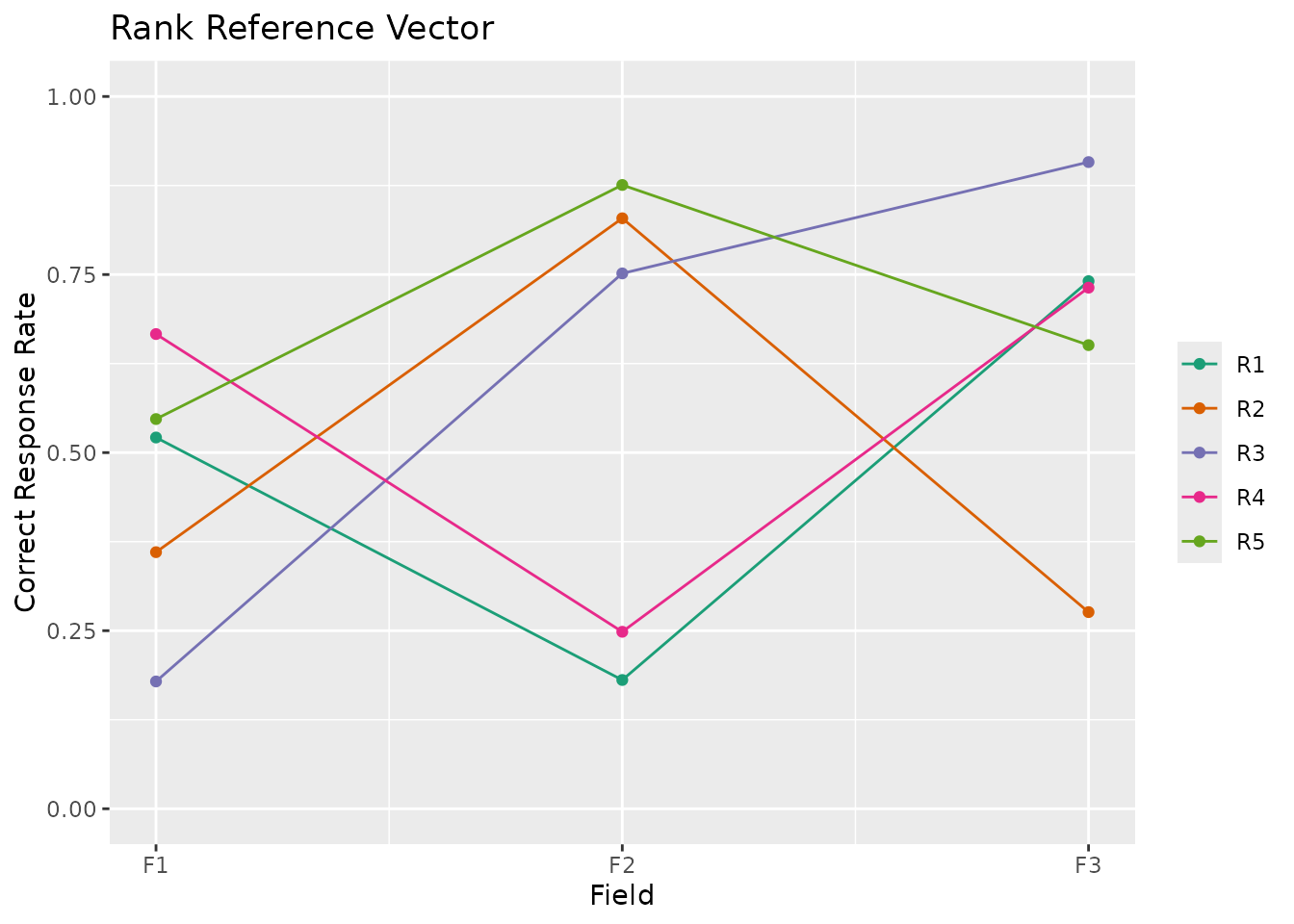 # \donttest{
# Ordinal biclustering (polytomous)
data(J35S500)
result_ord <- Biclustering(J35S500, ncls = 5, nfld = 5, method = "R")
#> Ranklustering is chosen.
#>
iter 1 log_lik -22710.5
#>
iter 2 log_lik -21311.9
#>
iter 3 log_lik -21002.5
#>
iter 4 log_lik -20945.8
#>
iter 5 log_lik -20932.3
#>
iter 6 log_lik -20929.2
#>
iter 7 log_lik -20929.8
plotRRV_gg(result_ord) # Default: mean
# \donttest{
# Ordinal biclustering (polytomous)
data(J35S500)
result_ord <- Biclustering(J35S500, ncls = 5, nfld = 5, method = "R")
#> Ranklustering is chosen.
#>
iter 1 log_lik -22710.5
#>
iter 2 log_lik -21311.9
#>
iter 3 log_lik -21002.5
#>
iter 4 log_lik -20945.8
#>
iter 5 log_lik -20932.3
#>
iter 6 log_lik -20929.2
#>
iter 7 log_lik -20929.8
plotRRV_gg(result_ord) # Default: mean
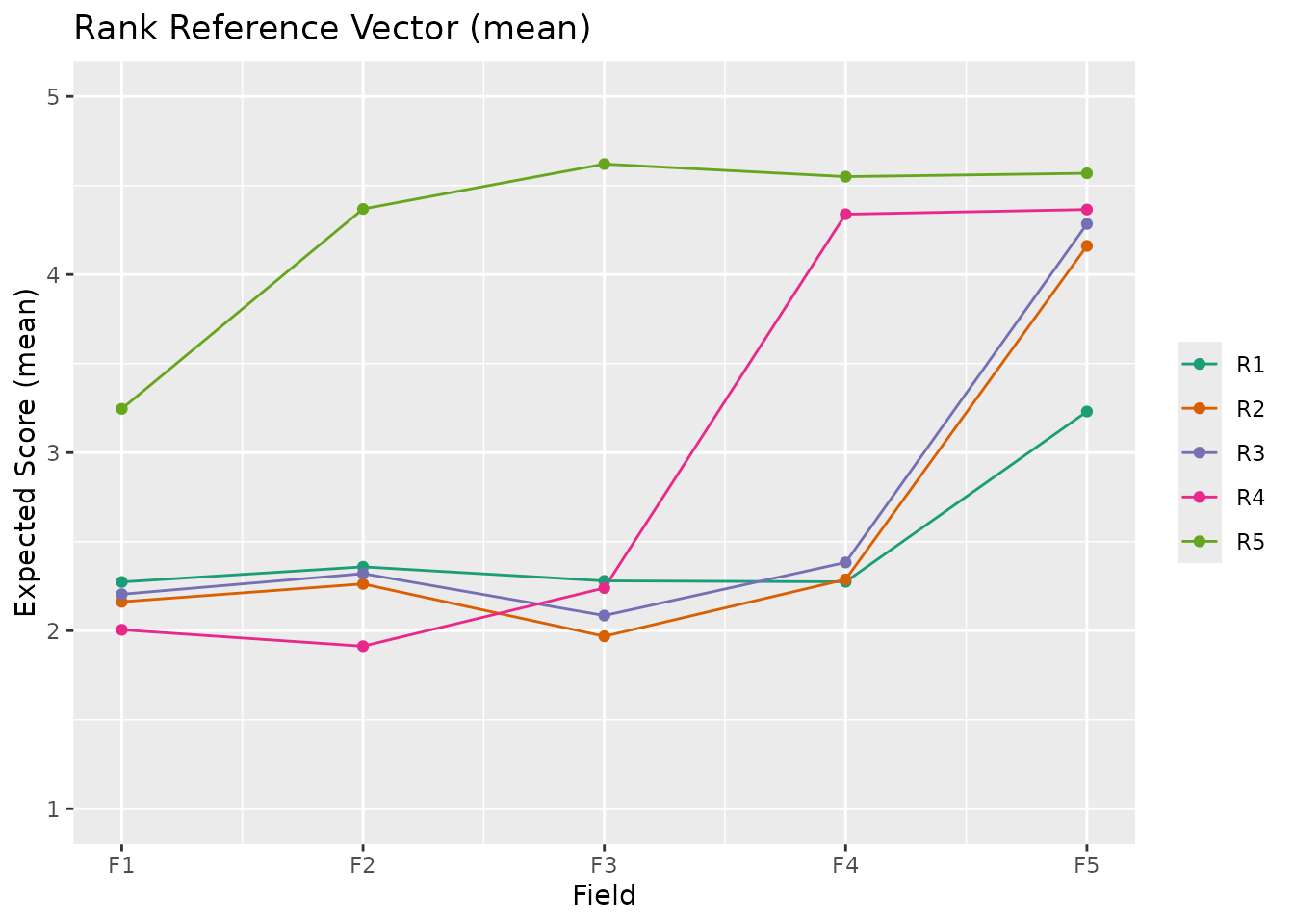 plotRRV_gg(result_ord, stat = "median")
plotRRV_gg(result_ord, stat = "median")
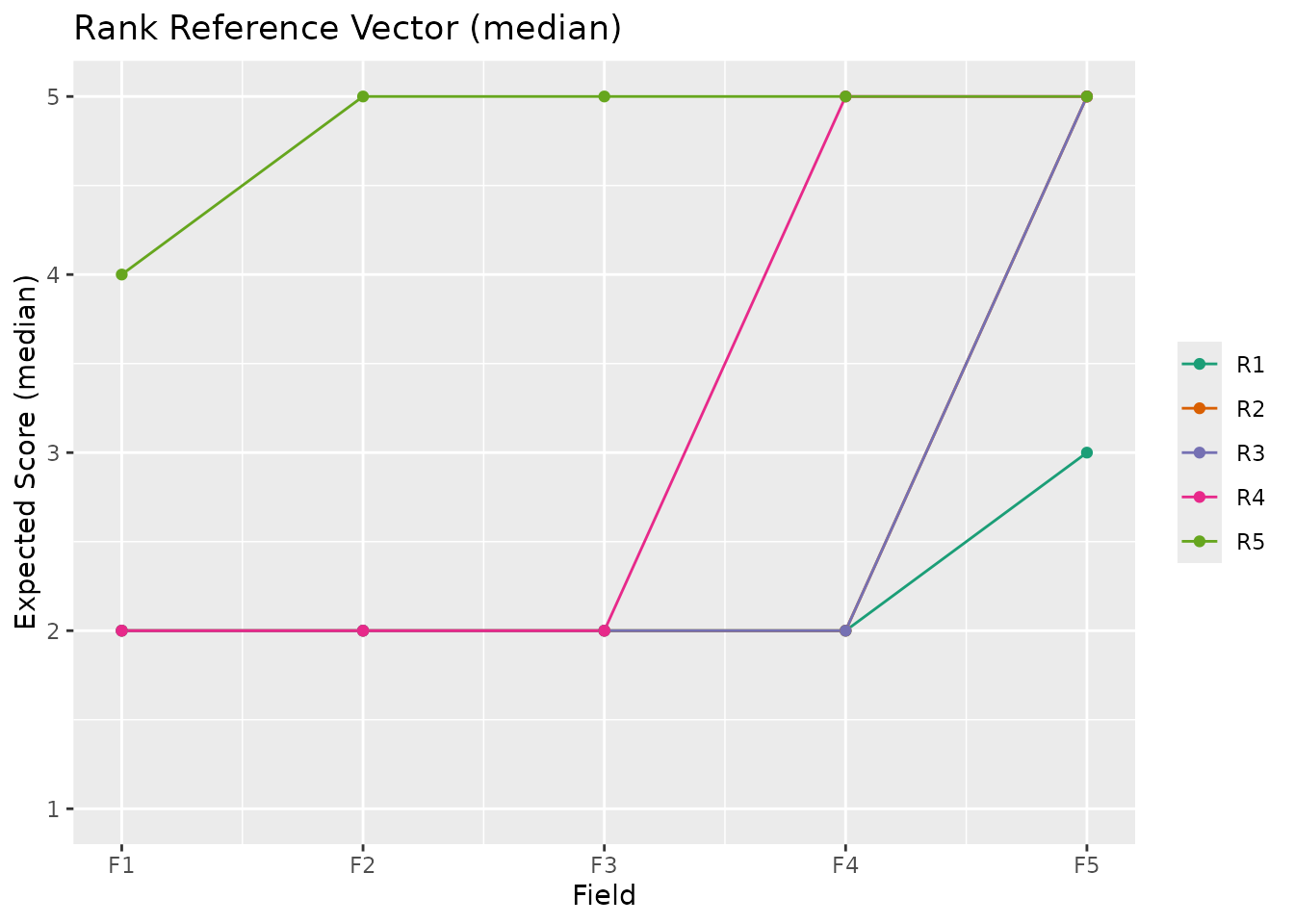 plotRRV_gg(result_ord, stat = "mode")
plotRRV_gg(result_ord, stat = "mode")
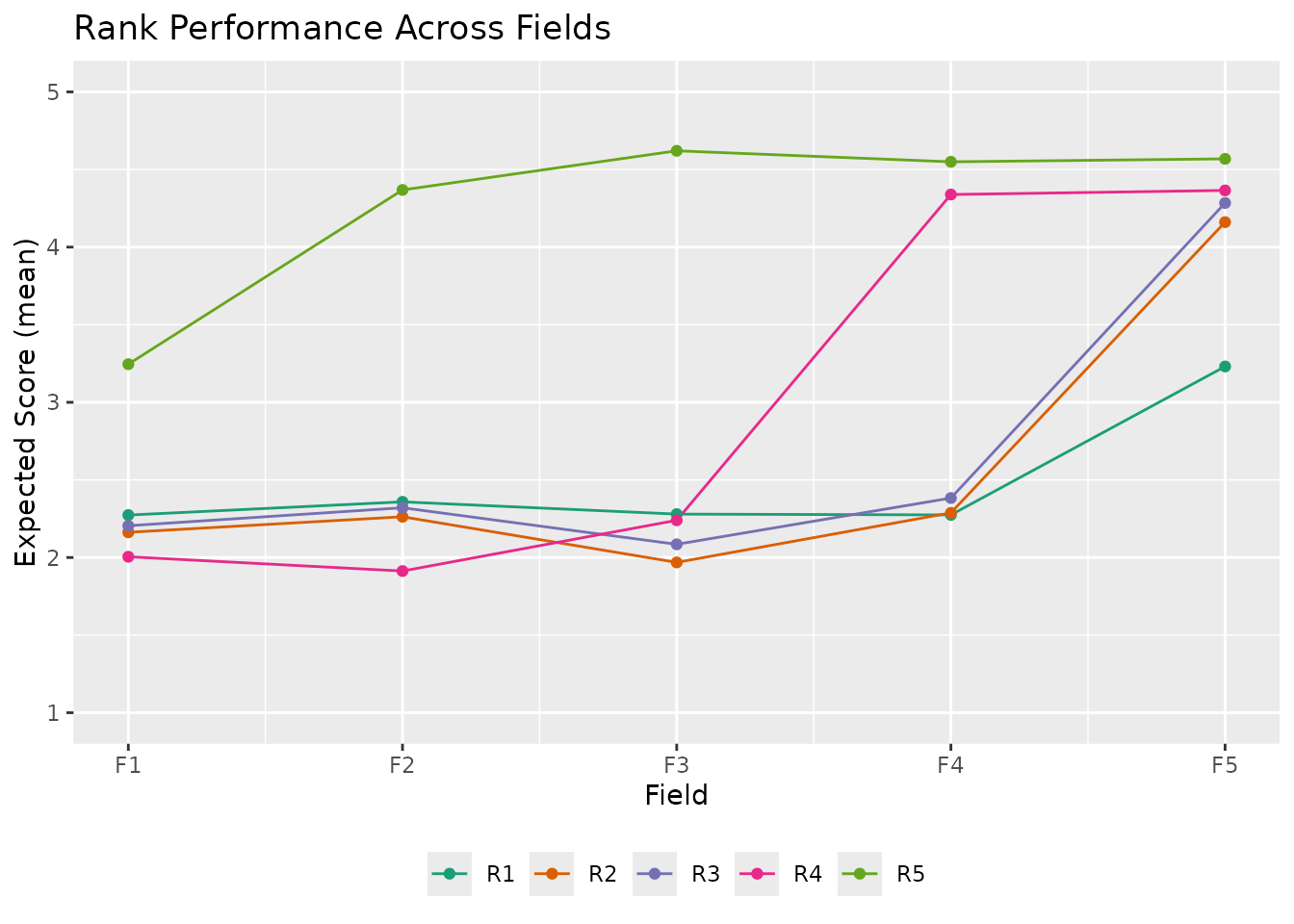
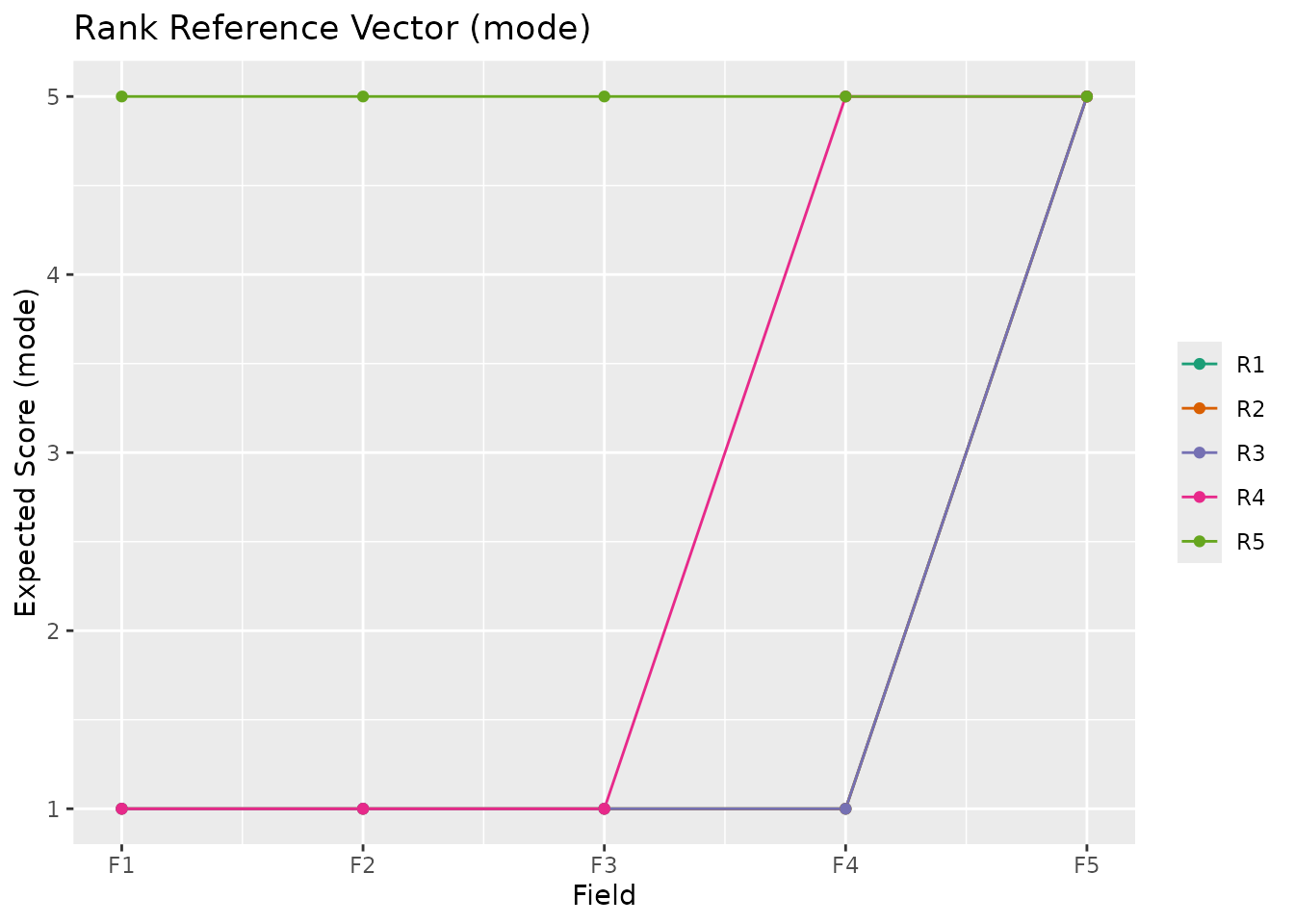 # Custom styling
plotRRV_gg(result_ord,
title = "Rank Performance Across Fields",
colors = c("#1b9e77", "#d95f02", "#7570b3", "#e7298a", "#66a61e"),
legend_position = "bottom"
)
# Custom styling
plotRRV_gg(result_ord,
title = "Rank Performance Across Fields",
colors = c("#1b9e77", "#d95f02", "#7570b3", "#e7298a", "#66a61e"),
legend_position = "bottom"
)
 # }
# }